Journal of Traditional Chinese Medicine ›› 2026, Vol. 46 ›› Issue (2): 490-500.DOI: 10.19852/j.cnki.jtcm.2026.02.020
• Original Articles • Previous Articles Next Articles
Candidate biomarker identification for blood stasis syndrome among coronary artery disease patients using the Olink proteomics platform
LI Hongzheng1,2, LIN Guosheng3, PENG Yuxuan1, Churov Alexey Viktorovich4, YANG Wenwen5, WANG Jie1, LU Jieming1, LIAO Feifei6, YU Ruotong1, WEI Yue1, ZHAO Zhiru1, LU Aimei6, LI Peng7, SHEN Aling3, LONG Linzi1(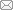 ), QU Hua1, FU Changgeng1(
), QU Hua1, FU Changgeng1( )
)
- 1
National Clinical Research Center for Chinese Medicine Cardiology ,Xiyuan Hospital, China Academy of Chinese Medical Sciences Beijing 100091, China
2Postdoctoral Research Center ,Guang’an Men Hospital, China Academy of Chinese Medical Sciences Beijing 100053, China
3Academy of Integrative Medicine ,Fujian University of Traditional Chinese Medicine Fuzhou 350122, China
4Institute of General Pathology and Pathophysiology ,Pirogov Russian National Research Medical University Moscow 125315, Russia
5Department of Cardiology ,Shaanxi Provincial Hospital of Chinese Medicine Xi’an 710003, China
6Academy of Chinese Medicine ,Beijing University of Chinese Medicine Beijing 100029, China
7Department of Cardiology I ,TCM Hospital Affiliated to Xinjiang Medical University Urumqi 830000, China
-
Received:2025-01-15Accepted:2025-07-02Online:2026-04-15Published:2026-04-04 -
Contact:Prof. FU Changgeng, National Clinical Research Center for Chinese Medicine Cardiology, Xiyuan Hospital, China Academy of Chinese Medical Sciences, Beijing 100091, China. fucgbs@163.com; Dr. LONG Linzi, National Clinical Research Center for Chinese Medicine Cardiology, Xiyuan Hospital, China Academy of Chinese Medical Sciences, Beijing 100091, China. qixiang830803@163.com; Telephone: +86-15101038490 -
Supported by:Formulation of International Diagnostic Criteria for Blood Stasis Syndrome in Coronary Heart Disease and Screening of Specific Protein Biomarkers(CMC2022010);Screening of Inflammatory Biomarkers and Analysis of Perivascular Adipose Tissue Characteristics in Patients with Blood Stasis Syndrome due to Coronary Heart Disease(GWJJMB202510021048);Comprehensive Investigation into Diagnostic Criteria and a Risk Prediction Model for Coronary Heart Disease with "Dryness-Blood Stasis Complex" Syndrome in Xinjiang(2022E02114)
Cite this article
LI Hongzheng, LIN Guosheng, PENG Yuxuan, Churov Alexey Viktorovich, YANG Wenwen, WANG Jie, LU Jieming, LIAO Feifei, YU Ruotong, WEI Yue, ZHAO Zhiru, LU Aimei, LI Peng, SHEN Aling, LONG Linzi, QU Hua, FU Changgeng. Candidate biomarker identification for blood stasis syndrome among coronary artery disease patients using the Olink proteomics platform[J]. Journal of Traditional Chinese Medicine, 2026, 46(2): 490-500.
share this article
| Item | CAD-BSS (n = 22) | CAD-non-BSS (n = 22) | Non-CAD-BSS (n = 22) | HC (n = 22) | P value |
|---|---|---|---|---|---|
| Age (years) | 61.32±15.88 | 71.05±7.69 | 52.05±11.45 | 54.91±11.04 | <0.001 |
| Gender (M/F, n) | 13/9 | 12/10 | 6/16 | 12/10 | - |
| BMI (kg/m²) | 24.47±2.91 | 23.35±3.08 | 23.46±2.70 | 25.44±3.99 | 0.292 |
| SBP (mm Hg) | 140.09±17.56 | 131.64±17.21 | 133.14±15.82 | 129.14±12.96 | 0.221 |
| DBP (mm Hg) | 78.59±9.79 | 78.59±10.93 | 84.32±13.91 | 84.86±8.45 | 0.106 |
| Hypertension [n (%)] | 17 (77.27) | 15 (68.18) | 5 (22.73) | 9 (40.91) | - |
| T2DM [n (%)] | 7 (31.82) | 10 (45.45) | 4 (18.18) | 1 (4.55) | - |
| Stroke [n (%)] | 2 (9.09) | 4 (18.18) | 0 (0.00) | 1 (4.55) | - |
| Hyperlipidemia [n (%)] | 3 (13.64) | 3 (13.64) | 1 (4.55) | 2 (9.09) | - |
| Antilaminate [n (%)] | 8 (36.36) | 5 (22.73) | 2 (9.09) | 1 (4.55) | - |
| TG (mmol/L) | 1.17±0.51 | 1.35±1.06 | 1.09±0.65 | 1.47±0.60 | 0.163 |
| TCHO (mmol/L) | 3.71±0.89 | 3.62±0.85 | 4.33±1.41 | 4.41±1.21 | 0.005 |
| LDL-C (mmol/L) | 2.24±0.83 | 2.18±0.89 | 2.90±0.94 | 2.98±0.96 | 0.003 |
| HDL-C (mmol/L) | 1.19±0.27 | 1.13±0.29 | 1.34±0.32 | 1.07±0.32 | 0.049 |
| ApoA-1 (mg/L) | 1.28±0.13 | 1.25±0.15 | 1.24±0.15 | 1.16±0.12 | 0.030 |
| ApoA (mg/dL) | 17.80±20.61 | 26.95±27.35 | 10.53±19.94 | 6.93±12.56 | <0.001 |
| PT (s) | 11.37±0.70 | 11.60±0.78 | 11.26±0.70 | 11.25±0.47 | 0.219 |
| PTT (%) | 101.95±6.79 | 99.81±7.80 | 103.00±6.96 | 103.14±5.17 | 0.219 |
| INR (%) | 0.96±0.06 | 0.98±0.07 | 0.95±0.06 | 0.95±0.04 | 0.239 |
| APTT (s) | 27.37±1.92 | 27.72±1.76 | 27.11±2.23 | 28.65±1.79 | 0.024 |
| FIB (g/L) | 3.13±0.74 | 3.16±0.86 | 3.62±3.27 | 2.88±0.58 | 0.461 |
| TT (s) | 16.86±1.23 | 18.54±12.48 | 16.01±3.37 | 17.60±0.89 | 0.007 |
| RBC (×1012/L) | 4.50±0.74 | 4.23±0.56 | 4.32±0.64 | 4.79±0.57 | 0.041 |
| PLT (×109/L) | 230.41±54.63 | 202.27±60.44 | 239.95±76.80 | 233.77±43.82 | 0.057 |
| HBG (g/L) | 135.23±16.41 | 131.50±17.65 | 130.91±21.09 | 147.91±15.61 | 0.015 |
| PCT (%) | 0.24±0.05 | 0.22±0.06 | 0.25±0.06 | 0.24±0.05 | 0.415 |
| WBC (×109/L) | 6.50±2.19 | 6.45±1.36 | 5.48±1.63 | 5.96±1.61 | 0.117 |
| NEUT (×109/L) | 4.44±1.61 | 4.29±1.35 | 3.36±1.33 | 3.51±1.34 | 0.027 |
| LYMP (×109/L) | 1.55±0.76 | 1.70±0.50 | 1.66±0.45 | 2.00±0.51 | 0.013 |
| Score of BSS (scores) | 3.50±1.41 | 0.00±0.00 | 3.59±1.62 | 0.00±0.00 | <0.001 |
Table 1 Basic clinical information of all patients ($ \bar{x} \pm s$)
| Item | CAD-BSS (n = 22) | CAD-non-BSS (n = 22) | Non-CAD-BSS (n = 22) | HC (n = 22) | P value |
|---|---|---|---|---|---|
| Age (years) | 61.32±15.88 | 71.05±7.69 | 52.05±11.45 | 54.91±11.04 | <0.001 |
| Gender (M/F, n) | 13/9 | 12/10 | 6/16 | 12/10 | - |
| BMI (kg/m²) | 24.47±2.91 | 23.35±3.08 | 23.46±2.70 | 25.44±3.99 | 0.292 |
| SBP (mm Hg) | 140.09±17.56 | 131.64±17.21 | 133.14±15.82 | 129.14±12.96 | 0.221 |
| DBP (mm Hg) | 78.59±9.79 | 78.59±10.93 | 84.32±13.91 | 84.86±8.45 | 0.106 |
| Hypertension [n (%)] | 17 (77.27) | 15 (68.18) | 5 (22.73) | 9 (40.91) | - |
| T2DM [n (%)] | 7 (31.82) | 10 (45.45) | 4 (18.18) | 1 (4.55) | - |
| Stroke [n (%)] | 2 (9.09) | 4 (18.18) | 0 (0.00) | 1 (4.55) | - |
| Hyperlipidemia [n (%)] | 3 (13.64) | 3 (13.64) | 1 (4.55) | 2 (9.09) | - |
| Antilaminate [n (%)] | 8 (36.36) | 5 (22.73) | 2 (9.09) | 1 (4.55) | - |
| TG (mmol/L) | 1.17±0.51 | 1.35±1.06 | 1.09±0.65 | 1.47±0.60 | 0.163 |
| TCHO (mmol/L) | 3.71±0.89 | 3.62±0.85 | 4.33±1.41 | 4.41±1.21 | 0.005 |
| LDL-C (mmol/L) | 2.24±0.83 | 2.18±0.89 | 2.90±0.94 | 2.98±0.96 | 0.003 |
| HDL-C (mmol/L) | 1.19±0.27 | 1.13±0.29 | 1.34±0.32 | 1.07±0.32 | 0.049 |
| ApoA-1 (mg/L) | 1.28±0.13 | 1.25±0.15 | 1.24±0.15 | 1.16±0.12 | 0.030 |
| ApoA (mg/dL) | 17.80±20.61 | 26.95±27.35 | 10.53±19.94 | 6.93±12.56 | <0.001 |
| PT (s) | 11.37±0.70 | 11.60±0.78 | 11.26±0.70 | 11.25±0.47 | 0.219 |
| PTT (%) | 101.95±6.79 | 99.81±7.80 | 103.00±6.96 | 103.14±5.17 | 0.219 |
| INR (%) | 0.96±0.06 | 0.98±0.07 | 0.95±0.06 | 0.95±0.04 | 0.239 |
| APTT (s) | 27.37±1.92 | 27.72±1.76 | 27.11±2.23 | 28.65±1.79 | 0.024 |
| FIB (g/L) | 3.13±0.74 | 3.16±0.86 | 3.62±3.27 | 2.88±0.58 | 0.461 |
| TT (s) | 16.86±1.23 | 18.54±12.48 | 16.01±3.37 | 17.60±0.89 | 0.007 |
| RBC (×1012/L) | 4.50±0.74 | 4.23±0.56 | 4.32±0.64 | 4.79±0.57 | 0.041 |
| PLT (×109/L) | 230.41±54.63 | 202.27±60.44 | 239.95±76.80 | 233.77±43.82 | 0.057 |
| HBG (g/L) | 135.23±16.41 | 131.50±17.65 | 130.91±21.09 | 147.91±15.61 | 0.015 |
| PCT (%) | 0.24±0.05 | 0.22±0.06 | 0.25±0.06 | 0.24±0.05 | 0.415 |
| WBC (×109/L) | 6.50±2.19 | 6.45±1.36 | 5.48±1.63 | 5.96±1.61 | 0.117 |
| NEUT (×109/L) | 4.44±1.61 | 4.29±1.35 | 3.36±1.33 | 3.51±1.34 | 0.027 |
| LYMP (×109/L) | 1.55±0.76 | 1.70±0.50 | 1.66±0.45 | 2.00±0.51 | 0.013 |
| Score of BSS (scores) | 3.50±1.41 | 0.00±0.00 | 3.59±1.62 | 0.00±0.00 | <0.001 |
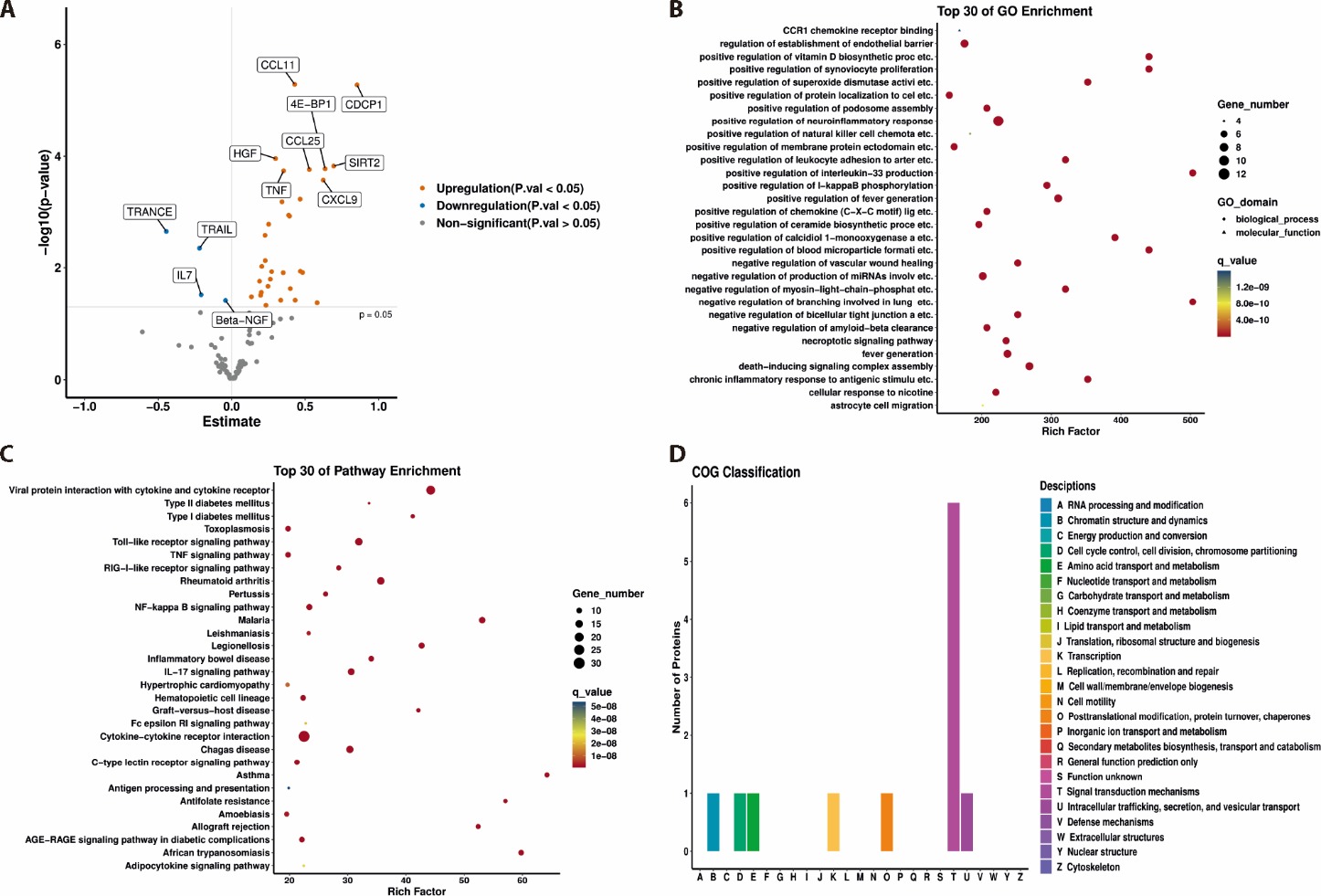
Figure 1 Identification of differentially expressed proteins in the CAD and non-CAD groups A: volcano plot of differentially expressed proteins in the CAD groups compared with the non-CAD groups; B: top 30 of GO enrichment; C: top 30 of pathway enrichment; D: COG classification of the differentially expressed proteins.CAD: coronary artery disease; GO: Geno Ontology; COG: clusters of orthologous groups; CCL: CC chemokine ligand; 4E-BP1: 4E-binding protein 1; CDCP1: CUB domain-containing protein 1; SIRT2: sirtuin 2; HGP: SLC25A16, solute carrier family 25 member 16; TNF: tumor necrosis factor; CXCL9: C-X-C motif chemokine ligand 9; TRAIL: TNF-related apoptosis-inducing ligand; TRANCE: TNF-related activation-induced cytokine; IL: interleukin; Beta-NGF: Beta nerve growth factor.
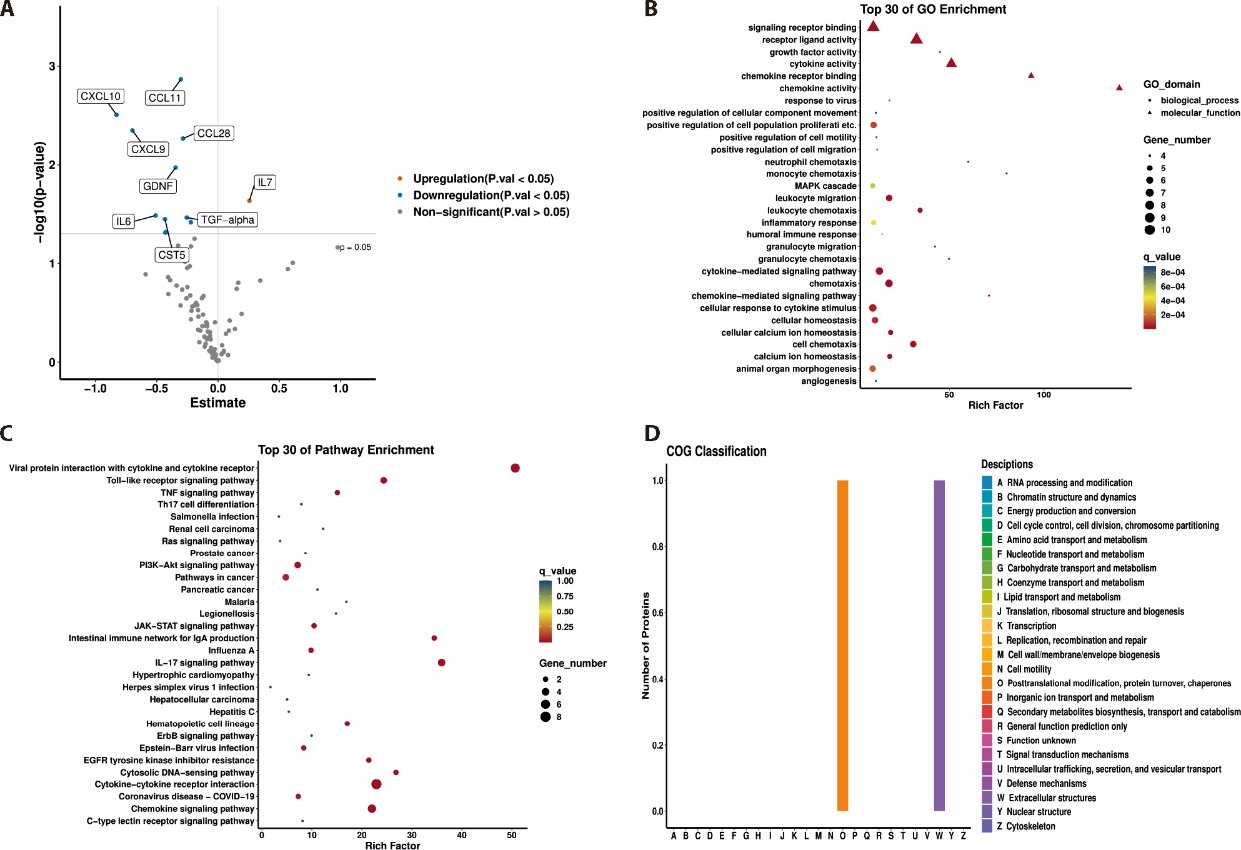
Figure 2 Identification of differentially expressed proteins in the CAD-BSS and CAD-non-BSS groups A: volcano plot of differentially expressed proteins in the CAD-BSS group compared with the CAD-non-BSS group; B: GO enrichment analysis of the differentially expressed proteins; C: pathway enrichment analysis of the differentially expressed proteins; D: COG classification of the differentially expressed protein. CAD: coronary artery disease; BSS: blood stasis syndrome; GO: Geno Ontology; COG: clusters of orthologous groups; CCL: CC chemokine ligand; CST5: cystatin D; STAMBP: STAM binding protein; SIRT2: sirtuin 2; MMP-10: matrix metalloproteinase 10; MCP-4: monocyte chemoattractant protein 4; IL: interleukin.
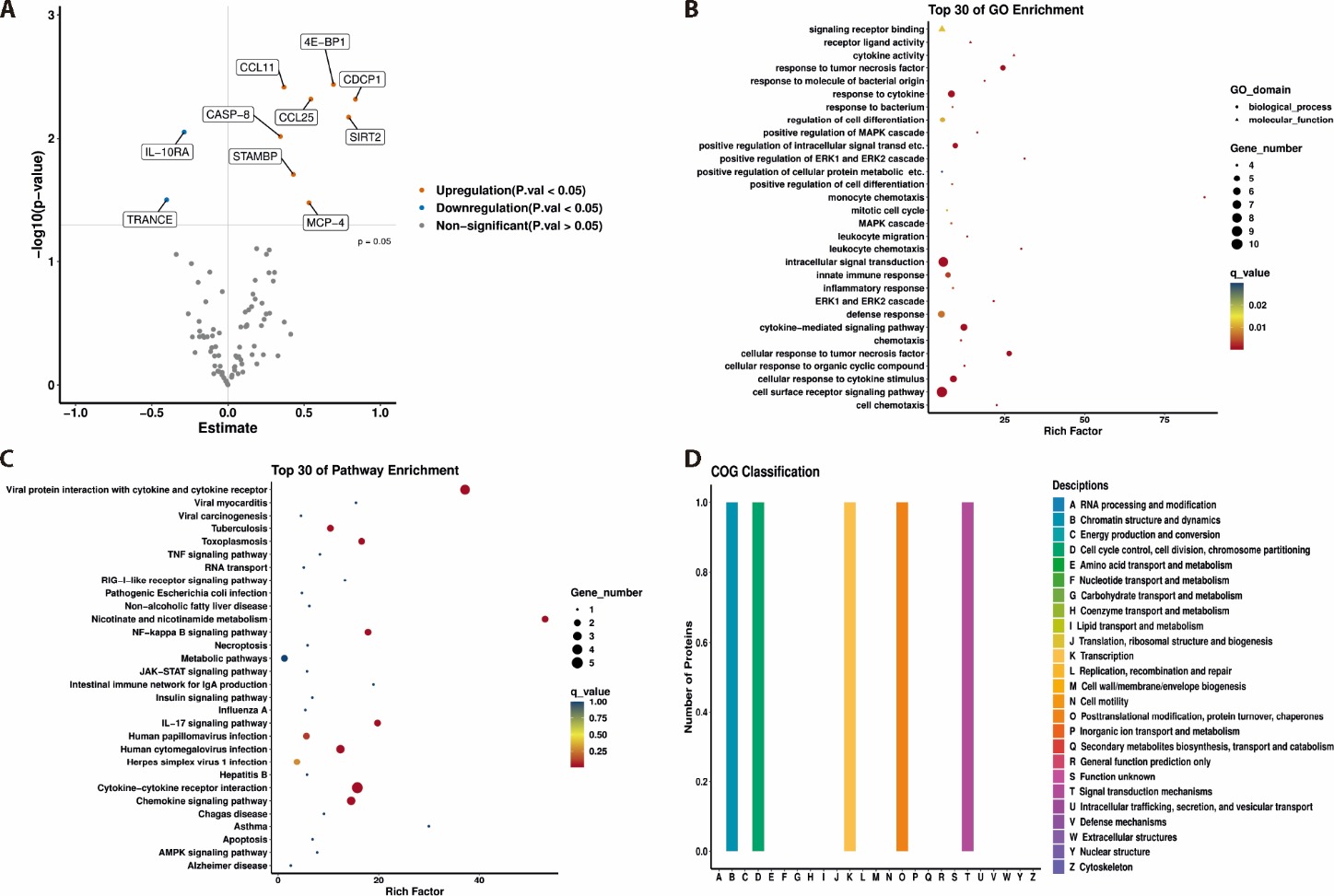
Figure 3 Identification of differentially expressed proteins in the CAD-BSS and non-CAD-BSS groups A: volcano plot of differentially expressed proteins in the CAD-BSS group compared with the non-CAD-BSS group; B: GO enrichment analysis of the differentially expressed proteins; C: pathway enrichment analysis of the differentially expressed proteins; D: COG classification of the differentially expressed protein. CAD: coronary artery disease; BSS: blood stasis syndrome; GO: Geno Ontology; COG: clusters of orthologous groups; 4E-BP1: 4E-binding protein 1; CASP-8: caspase-8; CCL: CC chemokine ligand; CDCP1: CUB domain-containing protein 1; IL-10RA: interleukin-10 receptor subunit alpha; MCP-4: monocyte chemoattractant protein 4; SIRT2: sirtuin 2; STAMBP: STAM binding protein; TRANCE: TNF-related activation-induced cytokine.
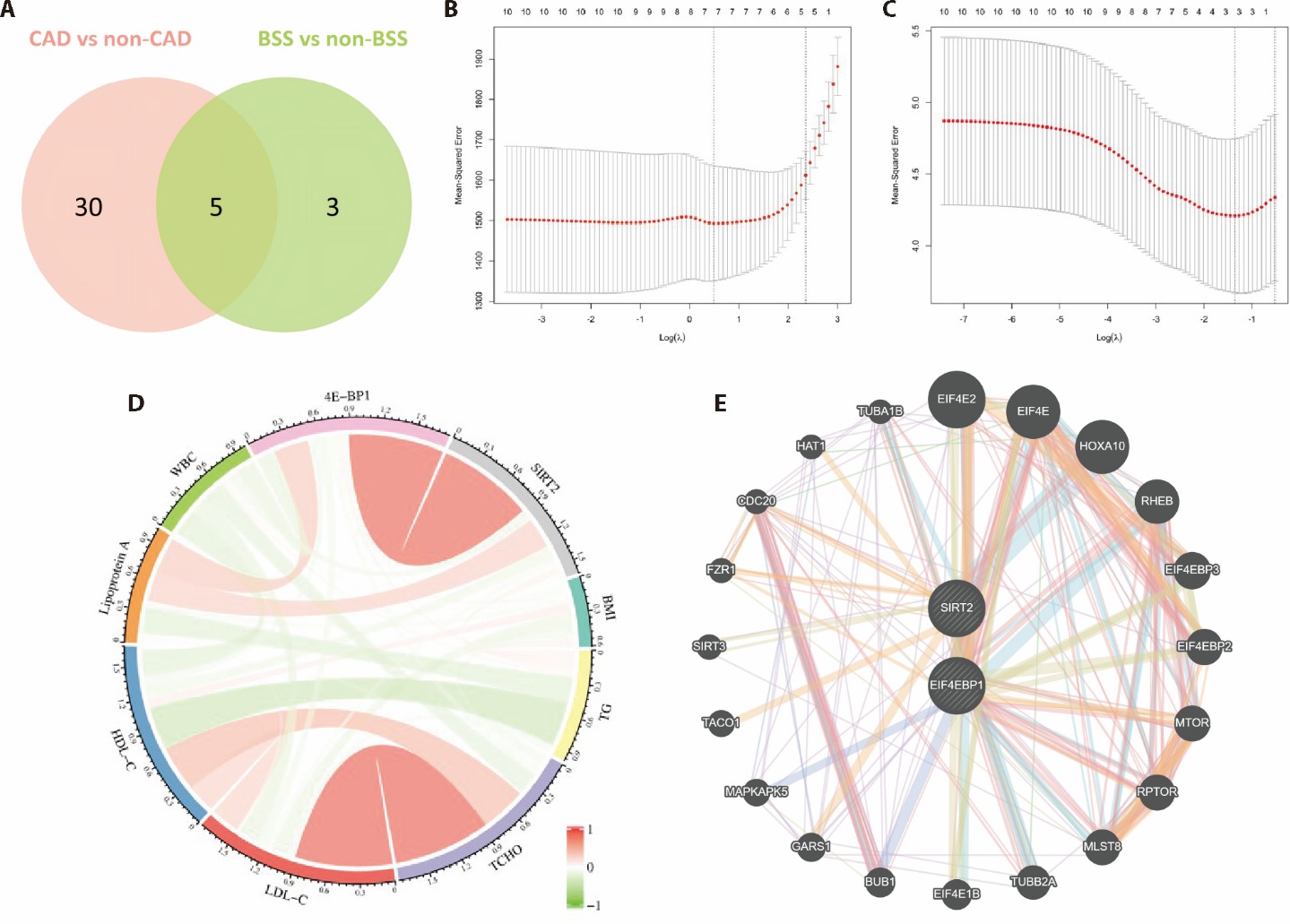
Figure 4 Identification of differentially expressed proteins between the HC, CAD-BSS, non-CAD-BSS, and CAD-non-BSS groups A: venn diagram of differentially expressed proteins in the four groups; B: LASSO regression analysis of the five proteins identified according to the presence/absence of a CAD diagnosis; C: LASSO regression analysis of the five proteins identified according to BSS scores; D: correlation analysis of the expression of the identified proteins with commonly used clinical indicators; E: the protein-protein interactional network of identified proteins, constructed according to the GeneMan database. CAD: coronary artery disease; BSS: blood stasis syndrome; 4E-BP1: 4e-binding protein 1; BMI: body mass index; BUB1: budding uninhibited by benzimidazoles 1; CDC20: cell division cycle 20; EIF4E: eukaryotic translation initiation factor 4e; EIF4EBP1: eukaryotic translation initiation factor 4e binding protein 1; FZR1: fizzy and cell division cycle 20 related 1; GARS1: glycyl-trna synthetase 1; HAT1: histone acetyltransferase 1; HDL-C: high-density lipoprotein cholesterol; HOXA10: homeobox a10; LDL-C: low-density lipoprotein cholesterol; MAPKAPK5: mapk activated protein kinase 5; MLST8: mtor associated protein, lst8 homolog; MTOR: mechanistic target of rapamycin; RHEB: ras homolog, mtorc1 binding; RPTOR: regulatory associated protein of mtor complex 1; SIRT: sirtuin; TACO1: translational activator of cytochrome c oxidase I; TCHO: total cholesterol; TG: triglyceride; TUBA1B: tubulin alpha 1b; TUBB2A: tubulin beta 2a class IIa; WBC: white blood cell count.
| 1. |
Konishi T, Virmani R, Jinnouchi H, et al. Plaque histological characteristics in individuals with sudden coronary death. Vascul Pharmacol 2023; 153: 107240.
DOI URL |
| 2. |
Yazdani AN, Pletsch M, Chorbajian A, Zitser D, Rai V, Agrawal DK. Biomarkers to monitor the prognosis, disease severity, and treatment efficacy in coronary artery disease. Expert Rev Cardiovasc Ther 2023; 21: 675-92.
DOI PMID |
| 3. |
Qiu Y, Xu H. Formulation and interpretation of international diagnostic guidelines for blood-stasis syndrome. Chin J Integr Med 2022; 28: 291-6.
DOI PMID |
| 4. |
Ma CY, Liu JH, Liu JX, et al. Relationship between two blood stasis syndromes and inflammatory factors in patients with acute coronary syndrome. Chin J Integr Med 2017; 23: 845-9.
DOI URL |
| 5. |
Li Y, Sun MQ, Li L, Zhang YH, Miao L, Liu JX. Application and prospect of platelet multi-omics technology in study of blood stasis syndrome. Chin J Integr Med 2022; 28: 99-105.
DOI |
| 6. |
Song M, Wang Y, Annex BH, Popel AS. Experiment-based computational model predicts that IL-6 classic and trans-signaling exhibit similar potency in inducing downstream signaling in endothelial cells. NPJ Syst Biol Appl 2023; 9: 45.
DOI PMID |
| 7. |
Ridker PM, Bhatt DL, Pradhan AD, et al. Inflammation and cholesterol as predictors of cardiovascular events among patients receiving statin therapy: a collaborative analysis of three randomised trials. Lancet 2023; 401: 1293-301.
DOI PMID |
| 8. |
Cardiovascular Professional Committee of the World Federation of Chinese Medicine Societies. International diagnostic guidelines for blood-stasis syndrome. Chin J Integr Med 2022; 28: 297-303.
DOI PMID |
| 9. | Otto CM, Nishimura RA, Bonow RO, et al. 2020 ACC/AHA guideline for the management of patients with valvular heart disease: executive summary: a report of the American College of Cardiology/American Heart Association Joint Committee on Clinical Practice Guidelines. Circulation 2021; 143: e35-71. |
| 10. |
Carlyle BC, Kitchen RR, Mattingly Z, et al. Technical performance evaluation of Olink proximity extension assay for blood-based biomarker discovery in longitudinal studies of Alzheimer's disease. Front Neurol 2022; 13: 889647.
DOI URL |
| 11. | Assarsson E, Lundberg M, Holmquist G, et al. Homogenous 96-plex PEA immunoassay exhibiting high sensitivity, specificity, and excellent scalability. PLoS One 2014; 9: e95192. |
| 12. |
Bardou P, Mariette J, Escudié F, Djemiel C, Klopp C. Jvenn: an interactive Venn diagram viewer. BMC Bioinformatics 2014; 15: 293.
DOI PMID |
| 13. |
Glickman JW, Dubin C, Renert-Yuval Y, et al. Cross-sectional study of blood biomarkers of patients with moderate to severe alopecia areata reveals systemic immune and cardiovascular biomarker dysregulation. J Am Acad Dermatol 2021; 84: 370-80.
DOI PMID |
| 14. |
Józefczuk E, Guzik TJ, Siedlinski M. Novel biomarkers and emerging tools to identify causal molecular pathways in hypertension and associated cardiovascular diseases. Kardiol Pol 2023; 81: 221-31.
DOI PMID |
| 15. |
Lind L, Ärnlöv J, Lindahl B, Siegbahn A, Sundström J, Ingelsson E. Use of a proximity extension assay proteomics chip to discover new biomarkers for human atherosclerosis. Atherosclerosis 2015; 242: 205-10.
DOI PMID |
| 16. |
Liu J, Chen B, Lu H, Chen Q, Li JC. Identification of novel candidate biomarkers for acute myocardial infarction by the Olink proteomics platform. Clin Chim Acta 2023; 548: 117506.
DOI URL |
| 17. |
Ponce-de-Leon M, Linseisen J, Peters A, et al. Novel associations between inflammation-related proteins and adiposity: a targeted proteomics approach across four population-based studies. Transl Res 2022; 242: 93-104.
DOI URL |
| 18. |
Benes J, Kroupova K, Kotrc M, et al. FGF-23 is a biomarker of RV dysfunction and congestion in patients with HFrEF. Sci Rep 2023; 13: 16004.
DOI |
| 19. |
Schmitz T, Wein B, Heier M, Peters A, Meisinger C, Linseisen J. Baseline fibroblast growth factor 23 is associated with long-term mortality in ST-elevation myocardial infarction-results from the Augsburg myocardial infarction registry. Front Cardiovasc Med 2023; 10: 1173281.
DOI URL |
| 20. |
Ueland GÅ, Methlie P, Heie A, et al. Substantial changes in inflammatory and cardiovascular biomarkers in patients with autonomous cortisol secretion. Eur J Endocrinol 2023; 189: 78-86.
DOI PMID |
| 21. |
Song X, Wang H, Wang C, et al. Association of sirtuin gene polymorphisms with susceptibility to coronary artery disease in a North Chinese Population. Biomed Res Int 2022; 2022: 4294008.
DOI URL |
| 22. | Yang W, Gao F, Zhang P, et al. Functional genetic variants within the SIRT2 gene promoter in acute myocardial infarction. PLoS One 2017; 12: e0176245. |
| 23. |
Demirci S, Doğan A, Apdik H, et al. Cytoglobin inhibits migration through PI3K/AKT/mTOR pathway in fibroblast cells. Mol Cell Biochem 2018; 437: 133-42.
DOI PMID |
| 24. |
Tait-Mulder J, Hodge K, Sumpton D, Zanivan S, Vazquez A. The conversion of formate into purines stimulates mTORC1 leading to CAD-dependent activation of pyrimidine synthesis. Cancer Metab 2020; 8:20.
DOI PMID |
| 25. |
Xin QQ, Chen X, Yuan R, et al. Correlation of platelet and coagulation function with blood stasis syndrome in coronary heart disease: a systematic review and Meta-analysis. Chin J Integr Med 2021; 27: 858-66.
DOI |
| 26. |
Wu G, Zhao J, Zhao J, et al. Exploring biological basis of Syndrome diffeentiation in coronary heart disease patients with two distinct Syndromes by integrated multi-omics and network pharmacology strategy. Chin Med 2021; 16: 109.
DOI |
| 27. |
Wang ZB, LI Y, Wang DP, et al. Proteomics analysis of coronary atherosclerotic heart disease with different Traditional Chinese Medicine syndrome types before and after percutaneous coronary intervention. J Tradit Chin Med 2024; 44: 554-63.
DOI |
| 28. |
Wang J, Liu L, Liu C, et al. Identification and analysis of differential miRNA-mRNA interactions in coronary heart disease: an experimental screening approach. Front Cardiovasc Med 2023; 10: 1186297.
DOI URL |
| 29. |
Mendes KL, Lelis DF, Santos SHS. Nuclear sirtuins and inflammatory signaling pathways. Cytokine Growth Factor Rev 2017; 38: 98-105.
DOI URL |
| 30. |
Zhang W, Liu D, Ren J, Zhou P, Han X. Overexpression of Sirtuin2 prevents high glucose-induced vascular endothelial cell injury by regulating the p53 and NF-κB signaling pathways. Biotechnol Lett 2018; 40: 271-8.
DOI PMID |
| 31. |
Guo J, Zhang H, Hu H, et al. Silent information regulator 2 deficiency exacerbates chronic cold exposure-induced colonic injury and p65 activation in mice. Gene 2024; 907: 148276.
DOI URL |
| 32. | Liu QQ, Ren K, Liu SH, Li WM, Huang CJ, Yang XH. MicroRNA-140-5p aggravates hypertension and oxidative stress of atherosclerosis via targeting Nrf2 and Sirt2. Int J Mol Med 2019; 43: 839-49. |
| 33. | Lum MA, Jonas KA, Parmar S, et al. Small-molecule modulators of B56-PP2A restore 4E-BP function to suppress eIF4E-dependent translation in cancer cells. J Clin Invest 2025; 135: e176093. |
| 34. |
Suginohara T, Wakabayashi K, Ato S, Ogasawara R. Effect of 2-deoxyglucose-mediated inhibition of glycolysis on the regulation of mTOR signaling and protein synthesis before and after high-intensity muscle contraction. Metabolism 2021; 114: 154419.
DOI URL |
| 35. | Chen T, Li T, Wang J. p 53 mediates PEDF-induced autophagy in human umbilical vein endothelial cells through sestrin2 signaling. Mol Med Rep 2019; 20: 1443-50. |
| 36. |
Böhm R, Imseng S, Jakob RP, Hall MN, Maier T, Hiller S. The dynamic mechanism of 4E-BP1 recognition and phosphorylation by mTORC1. Mol Cell 2021; 81: 2403-16.
DOI PMID |
| 37. |
Pi S, Mao L, Chen J, et al. The P2RY12 receptor promotes VSMC-derived foam cell formation by inhibiting autophagy in advanced atherosclerosis. Autophagy 2021; 17: 980-1000.
DOI URL |
| 38. |
Cao Y, Liu X, Lu W, et al. Fibronectin promotes cell proliferation and invasion through mTOR signaling pathway activation in gallbladder cancer. Cancer Lett 2015; 360:141-50.
DOI PMID |
| 39. |
Jeong YJ, Choi Y, Shin JM, et al. Melittin suppresses EGF-induced cell motility and invasion by inhibiting PI3K/Akt/mTOR signaling pathway in breast cancer cells. Food Chem Toxicol 2014; 68: 218-25.
DOI URL |
| 40. | Han Z, Yan G, Jousma J, et al. Translational regulation of SND1 governs endothelial homeostasis during stress. J Clin Invest 2025; 135: e168730. |
| 41. | Zhao YM, Zhang YL, Wang GZ, et al. Network pharmacology-based analysis of the antithrombotic clinical efficacy and antithrombotic mechanism of Huoxue Jiedu prescription in the treatment of polycythemia vera with heat toxin and blood stasis syndrome. J Tradit Chin Med 2025; 45: 1353-65. |
| [1] | XU Pingyuan, ZHU Heng, ZHOU Ruonan, WEI Yaping, ZHU Ziwei, XU Fangyuan, XIANG Yingying, CAO Yue, SHEN Lixuan, WANG Ziwei, XUE Yingying, YU Xizhong, FANG Penghua, SHANG Wenbin. Beneficial effects of Huanglian Jiedu decoction (黄连解毒汤 ) on metabolic syndrome: a prospective randomized controlled clinical trial [J]. Journal of Traditional Chinese Medicine, 2026, 46(2): 411-417. |
| [2] | YU Fangning, LIN Li, TANG Xiao, HE Xiujuan, ZHANG Bo, CHEN Yukun, MA Yizhao, LIU Zeyu, YE Jinsheng, XU Xuying. Effect of Huiyang Shengji unguent (回阳生肌膏) on the NOD-like receptor family pyrin domain containing protein 3/caspase1/ gasdermin D pathway and lymphatic angiogenesis in patients with diabetic foot ulcer [J]. Journal of Traditional Chinese Medicine, 2026, 46(2): 427-438. |
| [3] | GONG Yifan, XU Xiaohan, MA Jie, PANG Bo, LIU Wanlin, LIU Hongxiao. Multi-omics-based study on the biological characteristics of kidney renal deficiency and blood stasis in ankylosing spondylitis [J]. Journal of Traditional Chinese Medicine, 2026, 46(1): 183-194. |
| [4] | LI Dongqi, WANG Tongxing, WANG Zixuan, YAN Yihui, LI Jie, GU Jiaojiao, LI Cuiru, WANG Aili, SUN Lingling, MENG Yongjie, ZHANG Zeyu, HOU Yunlong, GAO Huailin. Improving glucose tolerance in obese rats: the role of Jinlida granules (津力达颗粒 ) in gut microbiota modulation [J]. Journal of Traditional Chinese Medicine, 2026, 46(1): 62-72. |
| [5] | WEI Mumu, YAN Xiaoyu, ZHANG Yujing, ZHANG Xinxin, YAN Yongbin. Soufeng Yuchuan formula (搜风愈喘方) alleviates asthma airway inflammation and suppresses the progression of asthma by inhibiting ferroptosis in airway epithelial cells [J]. Journal of Traditional Chinese Medicine, 2026, 46(1): 73-84. |
| [6] | ZOU Xiaoyun, WEI Minmin, JIA Shouning, QI Yongfu, LI Hailing, LIU Yan, SHI Xiujuan, LI Junru. Weimishu prescription (胃糜舒方) alleviates liver-stomach disharmony and chronic gastritis by reducing inflammation and regulating gastric stem cell differentiation [J]. Journal of Traditional Chinese Medicine, 2026, 46(1): 95-109. |
| [7] | LEI Xiaochun, LIU Cuizhen, LIN Xiujuan, XIE Xiangyu, KE Wei, QIU Zhenwen, TANG Hongmei, HUANG Yushen, ZHANG Lijuan, HUANG Baoyuan, WAN Xin, LI Detang. Effect of improved Yupingfeng powder prescription (玉屏风散加味) on interleukin-33/suppression of tumorigenicity 2 pathway in mice with ovalbumins-induced allergic rhinitis [J]. Journal of Traditional Chinese Medicine, 2025, 45(6): 1215-1227. |
| [8] | LI Keyao, SHU Ye, CHANG Jing, TANG Jianping, ZHANG Litao, WEI Zhu. Psoriasis intervention by Huai’er (Trametes): unveiling novel targets via network pharmacology [J]. Journal of Traditional Chinese Medicine, 2025, 45(6): 1317-1329. |
| [9] | WANG Wei, LONG Qi, FU Ling, WU Haiqiao. Jianpi Yifei Tongluo recipe (健脾益肺通络方剂) attenuates inflammation by promoting the expression of interferon regulatory factor 4 in the rat model of chronic obstructive pulmonary disease [J]. Journal of Traditional Chinese Medicine, 2025, 45(5): 1048-1058. |
| [10] | LI Weijia, LU Jing, MA Chao, LIU Mengmeng, PEI Ke, CHEN Hongyan, LIN Zhe, LYU Guangfu. Hamayou (Oviductus Ranae) protein hydrolysate ameliorates depression by regulating the mitogen-activated protein kinase pathway [J]. Journal of Traditional Chinese Medicine, 2025, 45(3): 493-507. |
| [11] | MIN Yu, ZHENG Meifeng, SUN Ju, PENG Zetong, CAO Zhixian, HUANG Xiaohua. Systematic acupuncture explains acupuncture at Baihui (GV20) and Fengchi (GB20) targeting the inflammatory response to regulate migraine [J]. Journal of Traditional Chinese Medicine, 2025, 45(3): 610-617. |
| [12] | YUAN Jiayao, WU Suhui, MENG Yufan, LI Hanbing, LI Genlin, XU Jiangyan. Yishen Tongluo formula (益肾通络方) ameliorates kidney injury via modulating inflammation and apoptosis in streptozotocin-induced diabetic kidney disease mice [J]. Journal of Traditional Chinese Medicine, 2025, 45(2): 254-265. |
| [13] | HUANG Jiaen, LUO Qing, DONG Gengting, PENG Weiwen, HE Jianhong, DAI Weibo. Xiahuo Pingwei San (夏藿平胃散) attenuated intestinal inflammation in dextran sulfate sodium-induced ulcerative colitis mice through inhibiting the receptor for advanced glycation end-products signaling pathway [J]. Journal of Traditional Chinese Medicine, 2025, 45(2): 311-325. |
| [14] | TIAN Yuan, BU He, WANG Tieshan, YANG Dongliang, ZHANG Wei, LIU Tong, ZHANG Li, HUO Zejun. Efficacy of electro-acupuncture at “Weizhong” (BL40) on macrophage polarization in rats with injured lumbar multifidus [J]. Journal of Traditional Chinese Medicine, 2025, 45(2): 335-347. |
| [15] | AN Yuanyuan, LIU Wang, LI Yanjie, WANG Yanchun, REN Xiaobin, HE Hongbing. Effect and mechanism of Sanqi (Radix Notoginseng) in treating periodontitis [J]. Journal of Traditional Chinese Medicine, 2025, 45(1): 66-75. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||

