Journal of Traditional Chinese Medicine ›› 2024, Vol. 44 ›› Issue (1): 122-130.DOI: 10.19852/j.cnki.jtcm.20231215.001
• Original articles • Previous Articles Next Articles
Efficacy of Ganshuang granules (肝爽颗粒) on non-alcoholic fatty liver and underlying mechanism: a network pharmacology and experimental verification
ZHI Guoguo1, SHAO Bingjie1, ZHENG Tianyan1, JI Shaoxiu1, LI Jingwei1, DANG Yanni2, LIU Feng2, WANG Dong1( )
)
- 1 School of Basic Medicine, Chengdu University of Traditional Chinese Medicine, Chengdu 611137, China
2 Shanxi Buchang Pharmaceutical Company, Limited, Xi’an 712000, China
-
Received:2022-12-22Accepted:2023-04-04Online:2024-02-15Published:2023-12-15 -
Contact:Dr. WANG Dong, School of Basic Medicine, Chengdu University of Traditional Chinese Medicine, Chengdu 611137, China. wangdong@cdutcm.edu.cn. Telephone: +86-15390022112 -
Supported by:Chengdu University of Traditional Chinese Medicine: Application of A Mouse Model to Explore the Mechanism of Therapeutic Action of Safranin in Type 2 Diabetes Mellitus(YXRC2018003)
Cite this article
ZHI Guoguo, SHAO Bingjie, ZHENG Tianyan, JI Shaoxiu, LI Jingwei, DANG Yanni, LIU Feng, WANG Dong. Efficacy of Ganshuang granules (肝爽颗粒) on non-alcoholic fatty liver and underlying mechanism: a network pharmacology and experimental verification[J]. Journal of Traditional Chinese Medicine, 2024, 44(1): 122-130.
share this article

Figure 1 PPI core target The smaller the degree, the lighter the color and the smaller the circle. PPI: protein-protein interaction; TNF: tumor necrosis factor; IL-6: interleukin 6; PIK3R1: phosphoinositide-3-kinase regulatory subunit 1; VEGFA: vascular endothelial growth factor A; CXCL8: C-X-C motif chemokine ligand 8; APP: amyloid precursor protein; STAT3: signal transducer and activator of transcription 3; Akt: protein kinase B; TP53: tumor protein p53.

Figure 2 Compound-target-pathway network diagram Green: circular nodes represent the active ingredients of GSG; purple: inverted triangle nodes represent signaling pathways; orange: rectangular nodes represent core goals. hsa04933: advanced glycation endproduct (AGE)-the receptor of advanced glycation endproduct (RAGE) signaling pathway in diabetic complications; hsa04657: IL-17 signaling pathway; hsa04066: hypoxia-inducible factor (HIF)-1 signaling pathway; hsa04920: adipocytokine signaling pathway; hsa04668: TNF signaling pathway; hsa04152: adenosine 5‘-monophosphate (AMP)-activated protein kinase (AMPK) signaling pathway; hsa04620: toll-like receptor signaling pathway; hsa04625: C-type lectin receptor signaling pathway; hsa04068: forkhead box O (FoxO) signaling pathway; hsa04151: PI3K-Akt signaling pathway; hsa04660: T cell receptor signaling pathway; hsa04910: insulin signaling pathway; hsa04630: Janus kinase (JAK)-STAT signaling pathway; hsa04010: mitogen-activated protein kinase (MAPK) signaling pathway; hsa04064: NF-kappa B signaling pathway; hsa04211: Longevity regulating pathway; hsa04621: NOD-like receptor signaling pathway; hsa04062: chemokine signaling pathway; hsa04115: tumor suppressor gene (p53) signaling pathway; hsa04370: VEGF signaling pathway; MOL007088: cryptotanshinone; MOL004635: saikosaponin A; MOL000422: kaempferol; MOL000098: quercetin; MOL000006: luteolin; MOL004328: naringenin; MOL007154: tanshinone IIA; MOL005828: nobiletin; MOL000472: emodin; MOL002268: rhein; MOL007074: salvianolic acid B; MOL005812: naringin; MOL005814: tangeretin; MOL001924: paeoniflorin; MOL000358: beta-sitosterol; MOL000354: isorhamnetin; MOL000043: atractylenolide; MOL013289: polydatin.
| Item | Control (n = 6) | Model (n = 6) | GSG (n = 6) |
|---|---|---|---|
| ALT (U/L) | 72.55±33.26 | 152.65±8.67a | 104.04±46.48b |
| AST (U/L) | 166.98±19.03 | 350.22±58.19a | 269.08±50.45b |
| ALP (U/L) | 102.10±6.23 | 138.03±24.10a | 103.56±23.21b |
| TC (mmol/L) | 3.62±0.57 | 5.07±0.44a | 3.58±0.89b |
| TG (mmol/L) | 0.99±0.21 | 2.04±0.29a | 0.75±0.17b |
| LDL (mmol/L) | 0.26±0.04 | 0.32±0.04a | 0.26±0.06b |
Table 1 Changes in indicators related to liver injury and lipid metabolism were reflected in the serum of each group ($\bar{x}±s$)
| Item | Control (n = 6) | Model (n = 6) | GSG (n = 6) |
|---|---|---|---|
| ALT (U/L) | 72.55±33.26 | 152.65±8.67a | 104.04±46.48b |
| AST (U/L) | 166.98±19.03 | 350.22±58.19a | 269.08±50.45b |
| ALP (U/L) | 102.10±6.23 | 138.03±24.10a | 103.56±23.21b |
| TC (mmol/L) | 3.62±0.57 | 5.07±0.44a | 3.58±0.89b |
| TG (mmol/L) | 0.99±0.21 | 2.04±0.29a | 0.75±0.17b |
| LDL (mmol/L) | 0.26±0.04 | 0.32±0.04a | 0.26±0.06b |
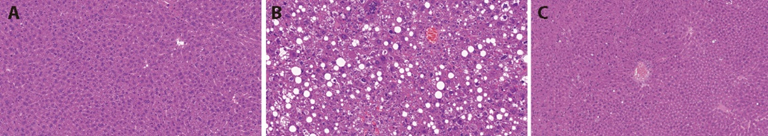
Figure 3 Results of liver pathological sections in each group by HE staining (× 20) A: control group; B: model group (× 20); C: GSG group (× 20). Control group: treated with normal diet and physiological saline; Model group: treated with high-fat diet and physiological saline; GSG group: treated with HFD and GSG of 0.1 mL/10 g. HE: hematoxylin and eosin; HFD: high-fat diet; GSG: Ganshuang granules.
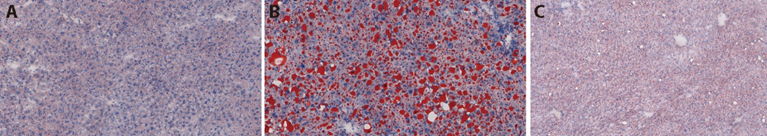
Figure 4 Expression of lipids in the liver of each group by oil red staining (× 20) A: control group; B: model group; C: GSG group. Control group: treated with normal diet and physiological saline; Model group: treated with high-fat diet and physiological saline; GSG group: treated with HFD and GSG of 0.1 mL/10 g. HFD: high-fat diet; GSG: Ganshuang granules.
| Item | Control (n = 6) | Model (n = 6) | GSG (n = 6) |
|---|---|---|---|
| LW (g) | 0.943±0.050 | 1.970±0.084a | 1.598±0.151b |
| BW (g) | 27.517±1.848 | 31.183±2.882a | 29.180±1.778b |
| LW/BW | 0.034±0.001 | 0.063±0.004a | 0.055±0.005b |
Table 2 Changes in mouse LW, BW, and LW/BW in each group ($\bar{x}±s$)
| Item | Control (n = 6) | Model (n = 6) | GSG (n = 6) |
|---|---|---|---|
| LW (g) | 0.943±0.050 | 1.970±0.084a | 1.598±0.151b |
| BW (g) | 27.517±1.848 | 31.183±2.882a | 29.180±1.778b |
| LW/BW | 0.034±0.001 | 0.063±0.004a | 0.055±0.005b |
| Item | Control (n = 4) | Model (n = 4) | GSG (n = 4) |
|---|---|---|---|
| IL-6 (pg/mg) | 181.912±27.074 | 299.723±85.682a | 206.379±32.358b |
| TNF-α (pg/mg) | 11.700±2.369 | 34.001±7.657a | 14.192±3.424b |
| SREBP-1/GAPDH | 0.431±0.081 | 0.847±0.032a | 0.643±0.020b |
| P-PI3K/GAPDH | 0.317±0.050 | 0.221±0.044a | 0.365±0.051b |
| P-AKT/GAPDH | 0.894±0.160 | 0.542±0.121a | 0.849±0.038b |
Table 3 Expression of proteins in P-PI3K, P-AKT, SREBP-1, IL-6 and TNF-α in liver tissues changed ($\bar{x}±s$)
| Item | Control (n = 4) | Model (n = 4) | GSG (n = 4) |
|---|---|---|---|
| IL-6 (pg/mg) | 181.912±27.074 | 299.723±85.682a | 206.379±32.358b |
| TNF-α (pg/mg) | 11.700±2.369 | 34.001±7.657a | 14.192±3.424b |
| SREBP-1/GAPDH | 0.431±0.081 | 0.847±0.032a | 0.643±0.020b |
| P-PI3K/GAPDH | 0.317±0.050 | 0.221±0.044a | 0.365±0.051b |
| P-AKT/GAPDH | 0.894±0.160 | 0.542±0.121a | 0.849±0.038b |
| 1. |
Eslam M, Valenti L, Romeo S. Genetics and epigenetics of NAFLD and NASH: clinical impact. J Hepatol 2018; 68: 268-79.
DOI PMID |
| 2. |
Sodum N, Kumar G, Bojja SL, Kumar N, Rao CM. Epigenetics in NAFLD/NASH: targets and therapy. Pharmacol Res 2021; 167: 105484.
DOI URL |
| 3. |
Chen G, Ni Y, Nagata N, Xu L, Ota T. Micronutrient antioxidants and nonalcoholic fatty liver disease. Int J Mol Sci 2016; 17: 1379.
DOI URL |
| 4. |
Zeigerer A. NAFLD-a rising metabolic disease. Mol Metab 2021; 50: 101274.
DOI URL |
| 5. | Li J, Zou B, Yeo YH, et al. Prevalence, incidence, and outcome of non-alcoholic fatty liver disease in Asia, 1999-2019: a systematic review and meta-analysis. Lancet Gastroenterol Hepatol 2019; 4: 389. |
| 6. |
Buzzetti E, Pinzani M, Tsochatzis EA. The multiple-hit pathogenesis of non-alcoholic fatty liver disease (NAFLD). Metabolism 2016; 65: 1038-48.
DOI URL |
| 7. |
Cobbina E, Akhlaghi F. Non-alcoholic fatty liver disease (NAFLD)- pathogenesis, classification, and effect on drug metabolizing enzymes and transporters. Drug Metab Rev 2017; 49: 197-211.
DOI PMID |
| 8. |
Safari Z, Gérard P. The links between the gut microbiome and non-alcoholic fatty liver disease (NAFLD). Cell Mol Life Sci 2019; 76: 1541-58.
DOI PMID |
| 9. |
Lau LHS, Wong SH. Microbiota, obesity and NAFLD. Adv Exp Med Biol 2018; 1061: 111-25.
DOI PMID |
| 10. |
Day CP, James OF. Steatohepatitis: a tale of two "hits"? Gastroenterology 1998; 114: 842-5.
DOI PMID |
| 11. |
Chalasani N, Younossi Z, Lavine JE, et al. The diagnosis and management of nonalcoholic fatty liver disease: practice guidance from the American Association for the Study of Liver Diseases. Hepatology 2018; 67: 328-57.
DOI PMID |
| 12. |
Chen M, Xie Y, Gong S, et al. Traditional Chinese Medicine in the treatment of nonalcoholic steatohepatitis. Pharmacol Res 2021; 172: 105849.
DOI URL |
| 13. |
Yang JY, Tao DQ, Ma W, et al. Protective effects and mechanisms of Lizhong decoction against non-alcoholic fatty liver disease in a rat model. J Tradit Chin Med 2022; 42: 773-80.
DOI PMID |
| 14. |
Li XL, Sun FX, Shang ZM, Zhang YX, Li J, Zhang QX. Effect of Dangfei Liganning capsule on liver X receptor α/steroid regulatory element binding protein-1/fatty acid synthase signal pathway in rats with metabolic- associated fatty liver disease. J Tradit Chin Med 2022; 42: 940-7.
DOI PMID |
| 15. | Pang XC, Kang D, Fang JS, et al. Network pharmacology-based analysis of Chinese herbal Naodesheng formula for application to Alzheimer's disease. Chin J Nat Med 2018; 16: 53-62. |
| 16. |
Gou X, Gao Z, Yang Y, et al. State-target strategy: a bridge for the integration of Chinese and Western Medicine. J Tradit Chin Med 2021; 41: 1-5.
DOI PMID |
| 17. | Sun HQ, Wang XQ, Shi HB, Lou JL, Chen Y, Duan ZP. Ganshuang granules protect mouse liver from chronic injury induced by CCl4 via autophagy. Lin Chuang Gan Dan Bing Za Zhi 2015; 31: 1114-9. |
| 18. | Liu L LJ, Liu CY, et al. Effect of tenofovir combined with Ganshuang granule on hemodynamics of portal vein system in hepatitis B liver cirrhosis. Lin Chuang Gan Dan Bing Za Zhi 2021; 42: 134-9. |
| 19. | Lu F, Geng JB, Zhang JW, Dong Y. Effect of entecavir plus Ganshuang granule on fibrosis and cirrhosis in patients with chronic hepatitis B. J Tradit Chin Med 2021; 41: 624-9. |
| 20. |
Li S, Zhang B. Traditional Chinese Medicine network pharmacology: theory, methodology and application. Chin J Nat Med 2013; 11: 110-20.
DOI URL |
| 21. |
Kaixuan Z, Dong Z, Huiwei B, Lijing LI. Network pharmacology and molecular docking study on the effect of Kaempferol in treatment of metabolic associated fatty liver disease. J Tradit Chin Med 2022; 42: 788-94.
DOI PMID |
| 22. | Ru J, Li P, Wang J, et al. TCMSP: a database of systems pharmacology for drug discovery from herbal medicines. J Cheminform 2014; 6: 13. |
| 23. |
Huang J, Cheung F, Tan HY, et al. Identification of the active compounds and significant pathways of Yinchenhao decoction based on network pharmacology. Mol Med Rep 2017; 16: 4583-92.
DOI PMID |
| 24. |
Soudy M, Anwar AM, Ahmed EA, et al. UniprotR: retrieving and visualizing protein sequence and functional information from Universal Protein Resource (UniProt knowledgebase). J Proteomics 2020; 213: 103613.
DOI URL |
| 25. |
Li WH, Han JR, Ren PP, Xie Y, Jiang DY. Exploration of the mechanism of Zisheng Shenqi decoction against gout arthritis using network pharmacology. Comput Biol Chem 2021; 90: 107358.
DOI URL |
| 26. | Stelzer G, Rosen N, Plaschkes I, et al. The GeneCards Suite: from gene data mining to disease genome sequence analyses. Curr Protoc Bioinformatics 2016; 54: 1.30. 1-33. |
| 27. | Davis AP, Grondin CJ, Johnson RJ, et al. Comparative toxicogenomics database (CTD): update 2021. Nucleic Acids Res 2021; 49: D1138-43. |
| 28. | Amberger JS, Hamosh A. Searching online mendelian inheritance in man (OMIM): a knowledgebase of human genes and genetic phenotypes. Curr Protoc Bioinformatics 2017; 58: 1.2. 1-12. |
| 29. |
He D, Li Q, Du G, Meng G, Sun J, Chen S. An integration of network pharmacology and experimental verification to investigate the mechanism of Guizhi to treat nephrotic syndrome. Front Pharmacol 2021; 12: 755421.
DOI URL |
| 30. |
Zhang W, Zhang L, Wang WJ, et al. Network pharmacology and in vitro experimental verification to explore the mechanism of Sanhua decoction in the treatment of ischaemic stroke. Pharm Biol 2022; 60: 119-30.
DOI PMID |
| 31. | Wahab Khattak F, Salamah Alhwaiti Y, Ali A, Faisal M, Siddiqi MH. Protein-protein interaction analysis through network topology (Oral Cancer). J Healthc Eng 2021; 2021: 6623904. |
| 32. |
Shannon P, Markiel A, Ozier O, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 2003; 13: 2498-504.
DOI PMID |
| 33. |
Tang Y, Li M, Wang J, Pan Y, Wu FX. CytoNCA: a cytoscape plugin for centrality analysis and evaluation of protein interaction networks. Biosystems 2015; 127: 67-72.
DOI PMID |
| 34. |
Wan Y, Xu L, Liu Z, et al. Utilising network pharmacology to explore the underlying mechanism of Wumei pill in treating pancreatic neoplasms. BMC Complement Altern Med 2019; 19: 158.
DOI |
| 35. |
Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. Omics 2012; 16: 284-7.
DOI PMID |
| 36. |
Liu B, Deng X, Jiang Q, et al. Scoparone improves hepatic inflammation and autophagy in mice with nonalcoholic steatohepatitis by regulating the ROS/P38/Nrf 2 axis and PI3K/AKT/mTOR pathway in macrophages. Biomed Pharmacother 2020; 125: 109895.
DOI URL |
| 37. |
Zhao Q, Lin X, Wang G. Targeting SREBP-1-mediated lipogenesis as potential strategies for cancer. Front Oncol 2022; 12: 952371.
DOI URL |
| 38. |
Zeng S, Liu Y, Jiang C, Li B, Wen L, Feng Q. Clinical efficacy and safety of Ganshuang granules as an adjuvant treatment for chronic hepatitis B liver fibrosis: a protocol for systematic review and meta analysis. Medicine (Baltimore) 2020; 99: e22692.
DOI URL |
| 39. |
Zhang R, Zhu X, Bai H, Ning K. Network pharmacology databases for Traditional Chinese Medicine: review and assessment. Front Pharmacol 2019; 10: 123.
DOI PMID |
| 40. |
Van De Wier B, Koek GH, Bast A, Haenen GR. The potential of flavonoids in the treatment of non-alcoholic fatty liver disease. Crit Rev Food Sci Nutr 2017; 57: 834-55.
PMID |
| 41. |
Zhu Y, Liu R, Shen Z, Cai G. Combination of luteolin and lycopene effectively protect against the "two-hit" in NAFLD through Sirt1/AMPK signal pathway. Life Sci 2020; 256: 117990.
DOI URL |
| 42. |
Abu-Elsaad N, El-Karef A. Protection against nonalcoholic steatohepatitis through targeting IL-18 and IL-1 alpha by luteolin. Pharmacol Rep 2019; 71: 688-94.
DOI PMID |
| 43. |
Yin Y, Gao L, Lin H, et al. Luteolin improves non-alcoholic fatty liver disease in db/db mice by inhibition of liver X receptor activation to down-regulate expression of sterol regulatory element binding protein 1c. Biochem Biophys Res Commun 2017; 482: 720-6.
DOI URL |
| 44. |
Shanware NP, Bray K, Abraham RT. The PI3K, metabolic, and autophagy networks: interactive partners in cellular health and disease. Annu Rev Pharmacol Toxicol 2013; 53: 89-106.
DOI PMID |
| 45. |
Shaywitz AJ, Courtney KD, Patnaik A, Cantley LC. PI3K enters beta-testing. Cell Metab 2008; 8: 179-81.
DOI PMID |
| 46. | Matsuda S, Kobayashi M, Kitagishi Y. Roles for PI3K/AKT/PTEN pathway in cell signaling of nonalcoholic fatty liver disease. ISRN Endocrinol 2013; 2013: 472432. |
| 47. |
Huang X, Liu G, Guo J, Su Z. The PI3K/AKT pathway in obesity and type 2 diabetes. Int J Biol Sci 2018; 14: 1483-96.
DOI PMID |
| 48. |
Fan Y, He Z, Wang W, et al. Tangganjian decoction ameliorates type 2 diabetes mellitus and nonalcoholic fatty liver disease in rats by activating the IRS/PI3K/AKT signaling pathway. Biomed Pharmacother 2018; 106: 733-7.
DOI PMID |
| 49. |
Wang H, Tan H, Zhan W, et al. Molecular mechanism of Fufang Zhenzhu Tiaozhi capsule in the treatment of type 2 diabetes mellitus with nonalcoholic fatty liver disease based on network pharmacology and validation in minipigs. J Ethnopharmacol 2021; 274: 114056.
DOI URL |
| 50. |
Gu H, Jiang W, You N, et al. Soluble Klotho improves hepatic glucose and lipid homeostasis in type 2 diabetes. Mol Ther Methods Clin Dev 2020; 18: 811-23.
DOI URL |
| 51. |
Zeng L, Tang WJ, Yin JJ, Zhou BJ. Signal transductions and nonalcoholic fatty liver: a mini-review. Int J Clin Exp Med 2014; 7: 1624-31.
PMID |
| 52. |
Zhang J, Feng Q. Pharmacological effects and molecular protective mechanisms of astragalus polysaccharides on nonalcoholic fatty liver disease. Front Pharmacol 2022; 13: 854674.
DOI URL |
| 53. |
Chen Z, Yu R, Xiong Y, Du F, Zhu S. A vicious circle between insulin resistance and inflammation in nonalcoholic fatty liver disease. Lipids Health Dis 2017; 16: 203.
DOI PMID |
| 54. |
Risso G, Blaustein M, Pozzi B, Mammi P, Srebrow A. Akt/PKB: one kinase, many modifications. Biochem J 2015; 468: 203-14.
DOI PMID |
| 55. |
Jalali A, Zarshenas MM. AKT/GSK-3 pathway targeting; botanicals and bioactive compounds with anticancer activities. Curr Pharm Des 2021; 27: 3091-104.
DOI URL |
| 56. |
Revathidevi S, Munirajan AK. Akt in cancer: mediator and more. Semin Cancer Biol 2019; 59: 80-91.
DOI PMID |
| 57. |
Porta C, Paglino C, Mosca A. Targeting PI3K/Akt/mTOR signaling in cancer. Front Oncol 2014; 4: 64.
DOI PMID |
| 58. |
Hinz N, Jücker M. Distinct functions of AKT isoforms in breast cancer: a comprehensive review. Cell Commun Signal 2019; 17: 154.
DOI PMID |
| 59. | Jump DB. Fatty acid regulation of hepatic lipid metabolism. Curr Opin Clin Nutr Metab Care 2011; 14: 115-20. |
| 60. |
Xiao X, Song BL. SREBP: a novel therapeutic target. Acta Biochim Biophys Sin (Shanghai) 2013; 45: 2-10.
DOI URL |
| 61. |
Li L, Yang J, Li F, Gao F, Zhu L, Hao J. FBXW7 mediates high glucose-induced SREBP-1 expression in renal tubular cells of diabetic nephropathy under PI3K/Akt pathway regulation. Mol Med Rep 2021; 23: 233.
DOI URL |
| 62. |
Hao J, Liu S, Zhao S, et al. PI3K/Akt pathway mediates high glucose-induced lipogenesis and extracellular matrix accumulation in HKC cells through regulation of SREBP-1 and TGF-β1. Histochem Cell Biol 2011; 135: 173-81.
DOI PMID |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||

