Journal of Traditional Chinese Medicine ›› 2022, Vol. 42 ›› Issue (1): 116-121.DOI: 10.19852/j.cnki.jtcm.2022.01.008
• Research Articles • Previous Articles Next Articles
Efficacy of glucocorticoids, chloroquine and vitamin A on cytokine release syndrome: a network pharmacology study
Jing ZHANG, Jingjing ZHU, Siqi HE, Jianxun WANG(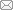 )
)
- School of Life Sciences, Beijing University of Chinese Medicine, Beijing, China.
-
Received:2021-03-03Accepted:2021-06-12Online:2022-02-15Published:2022-01-25 -
Contact:Jianxun WANG -
About author:Prof. WANG Jianxun, School of Life Sciences, Beijing University of Chinese Medicine, Beijing, China. Jianxun.Wang@bucm.edu.cn
-
Supported by:“Double-First Class” project of Beijing(1000041510155)
Cite this article
Jing ZHANG, Jingjing ZHU, Siqi HE, Jianxun WANG. Efficacy of glucocorticoids, chloroquine and vitamin A on cytokine release syndrome: a network pharmacology study[J]. Journal of Traditional Chinese Medicine, 2022, 42(1): 116-121.
share this article
| Gene | Mame | Primer (5'-3') |
|---|---|---|
| IL-2 | Forward | ACAGGATGCAACTCCTGTCT |
| Reverse | TGTGAGCATCCTGGTGAGTT | |
| IL-4 | Forward | CCAACTGCTTCCCCCTCTG |
| Reverse | TCTGTTACGGTCAACTCGGTG | |
| IL-10 | Forward | GCCAAGCCTTGTCTGAGATG |
| Reverse | GCATTCTTCACCTGCTCCAC | |
| TNF-α | Forward | AGCACTGAAAGCATGATCCG |
| Reverse | CCGATCACTCCAAAGTGCAG |
Table 1 Real-time reverse transcription-polymerase chain reaction primers sequences
| Gene | Mame | Primer (5'-3') |
|---|---|---|
| IL-2 | Forward | ACAGGATGCAACTCCTGTCT |
| Reverse | TGTGAGCATCCTGGTGAGTT | |
| IL-4 | Forward | CCAACTGCTTCCCCCTCTG |
| Reverse | TCTGTTACGGTCAACTCGGTG | |
| IL-10 | Forward | GCCAAGCCTTGTCTGAGATG |
| Reverse | GCATTCTTCACCTGCTCCAC | |
| TNF-α | Forward | AGCACTGAAAGCATGATCCG |
| Reverse | CCGATCACTCCAAAGTGCAG |
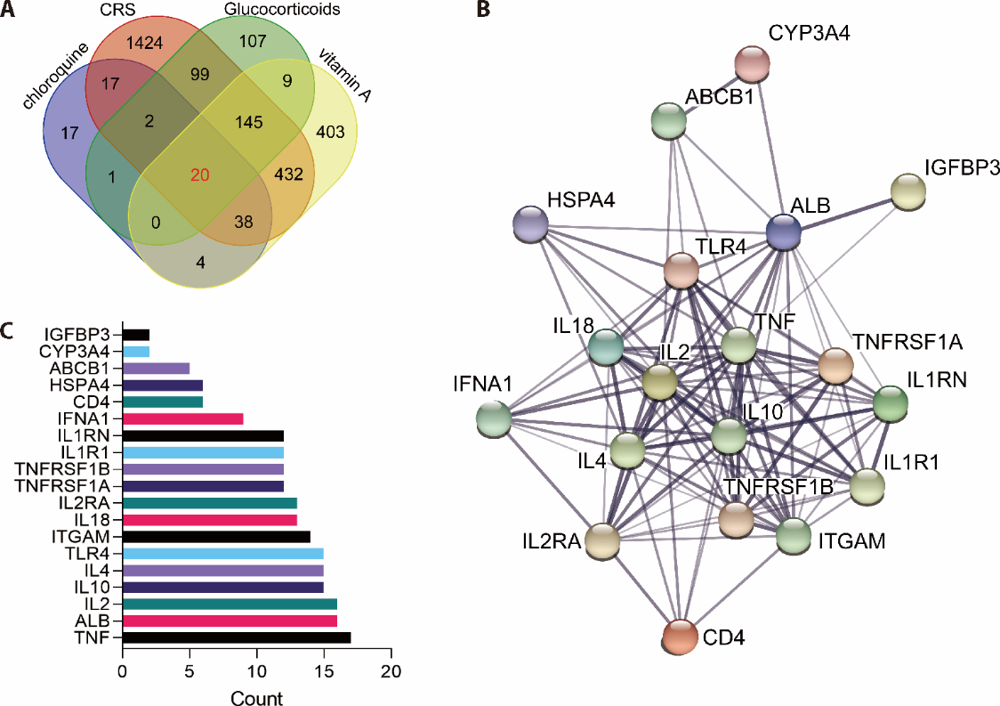
Figure 1 An association network of anti-inflammation drugs of glucocorticoids, chloroquine and vitamin A targeted genes associated with CRS A: Venn diagram of CRS and anti-inflammation drugs of glucocorticoids, chloroquine and vitamin A targeted genes; B: PPI network of 20 co-targeted genes was analyzed and calculated by STRING; C: the interaction scores of the 20 co-targeted genes in the PPI network were ranked. CYP3A4: cytochrome P450 family 3 subfamily A member 4; ABCB1: ATP binding cassette subfamily B member 1; HSPA4: heat shock protein family A member 4; ALB: albumin; IGFBP3: insulin like growth factor binding protein 3; TLR4: toll like receptor 4; IL-18: interleukin 18; IFNA1: interferon alpha 1; TNFRSF1A: TNF receptor superfamily member 1A; IL-1RN: interleukin 1 receptor antagonist; IL-1R1: interleukin 1 receptor type 1; TNFRSF1B: TNF receptor superfamily member 1B; IL-2RA: interleukin 2 receptor subunit alpha; ITGAM: integrin subunit alpha M; CRS: cytokine release syndrome.
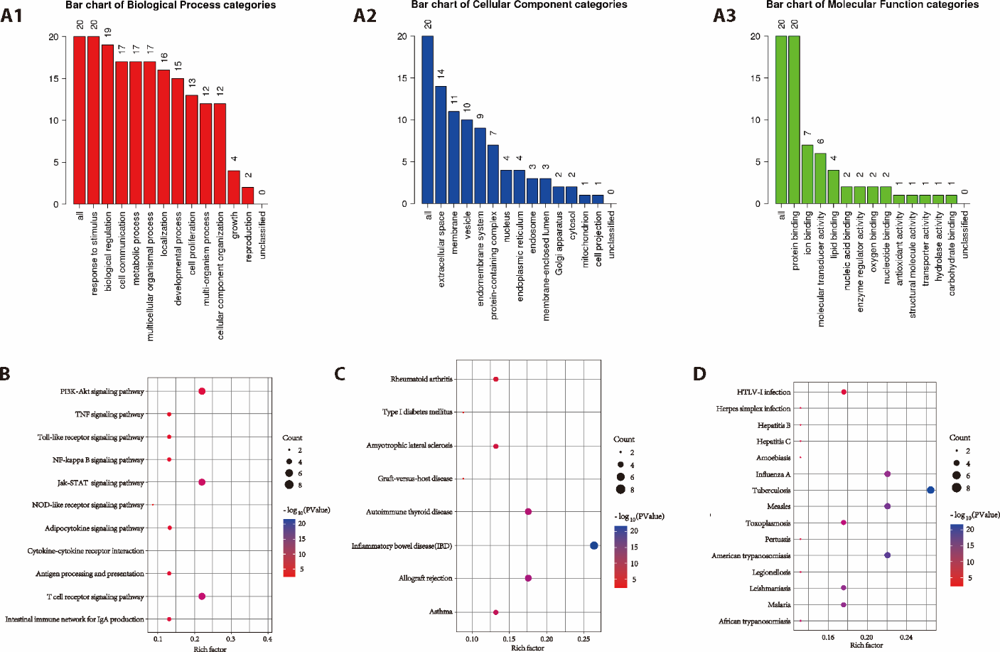
Figure 2 Gene ontology analysis and Kyoto Encyclopedia of Genes and Genomes pathway analysis A1-A3: the biological process, cell composition, and molecular function annotation of the 19 target genes; B: signal transduction pathway; C: non-infectious diseases associated pathway; D: infection associated pathway. The color scales indicate the different thresholds for the P-values, and the sizes of the dots represent the number of genes corresponding to each term.
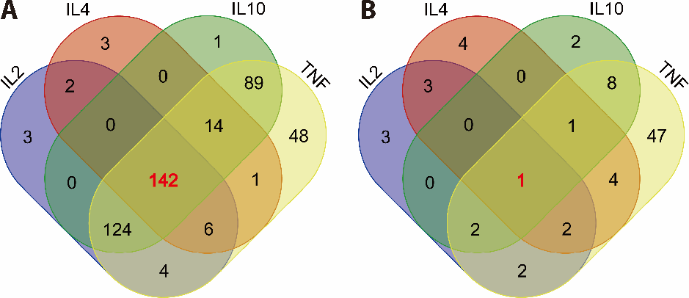
Figure 3 Venn diagram of drugs targeting IL-10, IL-4, IL-2 and TNF-α A: common ingredients targeting; B: Traditional Chinese Medicine targeting IL-10, IL-4, IL-2 and TNF-α. IL: interleukin; TNF-α: tumor necrosis factor-α.
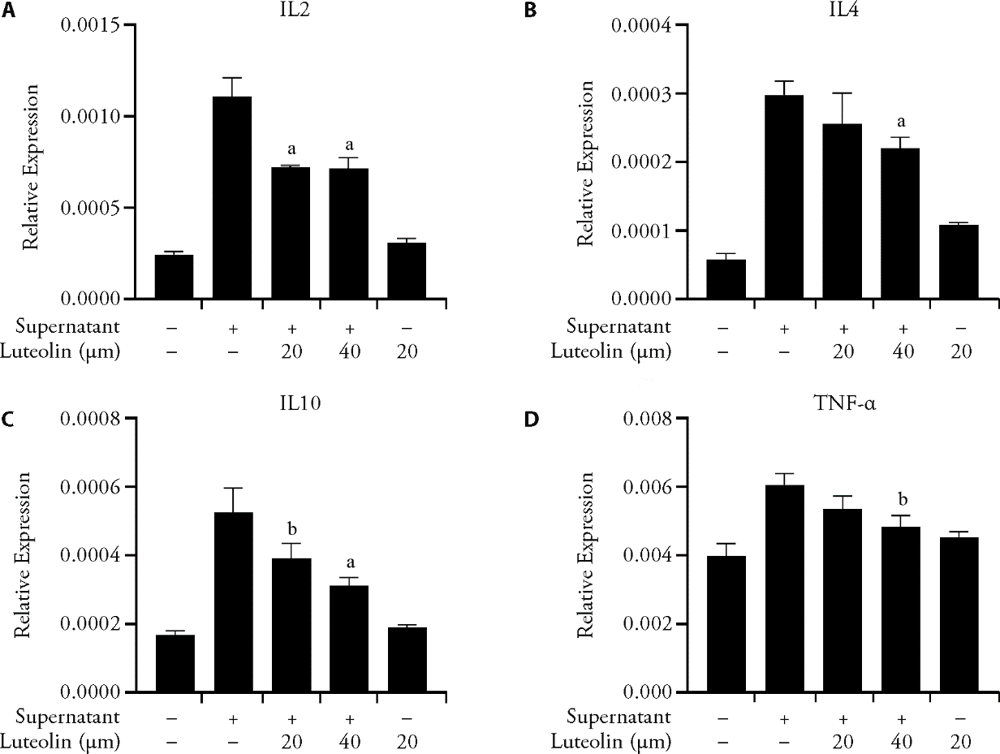
Figure 4 Regulation of expression of IL-10, IL-4, IL-2 and TNF-α in macrophages by Luteolin A: IL-2; B: IL-4; C: IL-10; D: TNF-α. Quantification of IL-10, IL-4, IL-2 and TNF-α mRNA was conducted by RT-PCR. The level of transcripts is normalized to β-actin mRNA. Error bars showed standard error of the mean, triplicate determinations. IL: interleukin; TNF-α: tumor necrosis factor-α. Statistical significance was calculated with analysis of variance with multiple comparisons, aP < 0.01, bP < 0.05.
| [1] | Teijaro JR. Cytokine storms in infectious diseases. Semin Immunopathol 2017;39:501-3. |
| [2] | Perrin P, Collongues N, Baloglu S, et al. Cytokine release syndrome-associated encephalopathy in patients with COVID-19. Eur J Neurol 2021;28:248-58. |
| [3] | Chousterman BG, Swirski FK, Weber GF. Cytokine storm and sepsis disease pathogenesis. Semin Immunopathol 2017;39:517-28. |
| [4] | Tisoncik JR, Korth MJ, Simmons CP, et al. Into the eye of the cytokine storm. Microbiol Mol Biol Rev 2012;76:16-32. |
| [5] | Lau SKP, Lau CCY, Chan KH, et al. Delayed induction of proinflammatory cytokines and suppression of innate antiviral response by the novel Middle East respiratory syndrome coronavirus: implications for pathogenesis and treatment. J Gen Virol 2013;94:2679-90. |
| [6] | Channappanavar R, Perlman S. Pathogenic human coronavirus infections: causes and consequences of cytokine storm and immunopathology. Semin Immunopathol 2017;39:529-39. |
| [7] | Hirano T, Murakami M. COVID-19: a new virus, but a familiar receptor and cytokine release syndrome. Immunity 2020;52:731-3. |
| [8] | Giavridis T, van der Stegen SJC, Eyquem J, et al. CAR T cell-induced cytokine release syndrome is mediated by macrophages and abated by IL-1 blockade. Nature Medicine 2018;24:731-8. |
| [9] | Hay KA. Cytokine release syndrome and neurotoxicity after CD19 chimeric antigen receptor-modified: CAR-) T cell therapy. Br J Haematol 2018;183:364-74. |
| [10] | Yao X, Huang JQ, Zhong HH, et al. Targeting interleukin-6 in inflammatory autoimmune diseases and cancers. Pharmacol Ther 2014;141:125-39. |
| [11] | Rossi JF, Lu ZY, Jourdan M, et al. Interleukin-6 as a therapeutic target. Clinical Cancer Research 2015;21:1248-57. |
| [12] | Colson P, Rolain JM, Raoult D. Chloroquine for the 2019 novel coronavirus SARS-CoV-2. Int J Antimicrob Agents 2020;55:105923. |
| [13] | Vincent MJ, Bergeron E, Benjannet S, et al. Chloroquine is a potent inhibitor of SARS coronavirus infection and spread. Virol J 2005;2. |
| [14] | Li R, Wu K, Li Y, et al. Revealing the targets and mechanisms of vitamin A in the treatment of COVID-19. Aging-Us 2020;12:15784-96. |
| [15] | Vandewalle J, Luypaert A, De Bosscher K, et al. Therapeutic mechanisms of glucocorticoids. Trends Endocrinol Metab 2018;29:42-54. |
| [16] | Chen Y, Yuan T, Chen D, et al. Systematic analysis of molecular mechanism of resveratrol for treating pulmonary hypertension based on network pharmacology technology. Eur J Pharmacol 2020;888:173466. |
| [17] | Kong Q, Wu Y, Gu Y, et al. Analysis of the molecular mechanism of Pudilan: PDL) treatment for COVID-19 by network pharmacology tools. Biomed Pharmacother 2020;128:110316. |
| [18] | Stelzer G, Rosen N, Plaschkes I, et al. The GeneCards suite: from gene data mining to disease genome sequence analyses. Curr Protoc Bioinformatics 2016; 54 54: 1.30.1-1.30.33. |
| [19] | Bateman A, Martin M-J, Orchard S, et al. UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Research 2021;49:D480-9. |
| [20] | Szklarczyk D, Gable AL, Lyon D, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Research 2019;47:D607-13. |
| [21] | Zhou YY, Zhou B, Pache L, et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nature Communications 2019;10:1523. |
| [22] | Ru JL, Li P, Wang JN, et al. TCMSP: a database of systems pharmacology for drug discovery from herbal medicines. J Cheminform 2014;6:13. |
| [23] | Ling CQ, Yue XQ, Ling C. Three advantages of using traditional Chinese medicine to prevent and treat tumor. J Integr Med 2014;12:331-5. |
| [24] | Brudno JN, Kochenderfer JN. Recent advances in CAR T-cell toxicity: mechanisms, manifestations and management. Blood Reviews 2019;34:45-55. |
| [25] | Caricchio R, Gallucci M, Dass C, et al. Preliminary predictive criteria for COVID-19 cytokine storm. Ann Rheum Dis 2021;80:88-95. |
| [26] | Qin YY, Zhou YH, Lu YQ, et al. Effectiveness of glucocorticoid therapy in patients with severe coronavirus disease 2019: protocol of a randomized controlled trial. Chin Med J: Engl) 2020;133:1080-6. |
| [27] | Pedersen SF, Ho YC. SARS-CoV-2: a storm is raging. J Clin Invest 2020;130:2202-5. |
| [28] | Han H, Ma QF, Li C, et al. Profiling serum cytokines in COVID-19 patients reveals IL-6 and IL-10 are disease severity predictors. Emerg Microbes Infect 2020;9:1123-30. |
| [29] | Riley JK, Takeda K, Akira S, et al. Interleukin-10 Receptor Signaling through the JAK-STAT Pathway. J Biol Chem 1999;274(23):16513-21. |
| [30] | Antoniv TT, Ivashkiv LB. Interleukin-10-induced gene expression and suppressive function are selectively modulated by the PI3K-Akt-GSK3 pathway. Immunology 2011;132:567-77. |
| [31] | Luo W, Li YX, Jiang LJ, et al. Targeting JAK-STAT Signaling to Control Cytokine Release Syndrome in COVID-19. Trends Pharmacol Sci 2020;41:531-43. |
| [32] | Nabavi SF, Braidy N, Gortzi O, et al. Luteolin as an anti-inflammatory and neuroprotective agent: a brief review. Brain Res Bull 2015;119:1-11. |
| [33] | A Xagorari AP, A Mauromatis, M Economou, et al. Luteolin inhibits an endotoxin-stimulated phosphorylation cascade and proinflammatory cytokine production in macrophages. J Pharmacol Exp Ther 2001;296:181-7. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||

