Journal of Traditional Chinese Medicine ›› 2022, Vol. 42 ›› Issue (6): 922-931.DOI: 10.19852/j.cnki.jtcm.20220817.002
• Research Articles • Previous Articles Next Articles
Transcriptome profile and its partly verification of human hepatocellular carcinoma cells exposed to Yuzhizi (Fructus Akebiae Quinatae) seed extract
FAN Xiaoyan1, LI Mingzhe1, CHEN Long2, LIANG Chao1, LU Wenli1, PAN Zhiqiang1, JIA Dongwei1, PENG Peike1, FANG Zhaoqin1( ), LIU Xiaomei1(
), LIU Xiaomei1( )
)
- 1 Department of Experimental Traditional Chinese Medicine, School of Basic Medical Science, Shanghai University of Traditional Chinese Medicine, Shanghai 201203, China
2 Experiment Center of Science and Technology, Shanghai University of Traditional Chinese Medicine, Shanghai 201203, China
-
Received:2021-11-11Accepted:2022-01-22Online:2022-12-15Published:2022-08-17 -
Contact:FANG Zhaoqin,LIU Xiaomei -
About author:LIU Xiaomei, Department of Experimental Traditional Chinese Medicine, School of Basic Medical Science, Shanghai University of Traditional Chinese Medicine, Shanghai 201203, China. wfjlxm2005@126.com, Telephone: +86-21-51322118; +86-21-51322117
FANG Zhaoqin, Department of Experimental Traditional Chinese Medicine, School of Basic Medical Science, Shanghai University of Traditional Chinese Medicine, Shanghai 201203, China. zq_fang@qq.com;
-
Supported by:grants from Shanghai Municipal Commission of Education: Study on the Synergistic Effect and Action Mechanism of Yuzhizi (Fructus Akebiae Quinatae) Seed Extract on Sorafenib(2019LK065);Shanghai Municipal Commission of Health and Family Planning [No. ](ZY (2018-2020)-CCCX-2001-01)
Cite this article
FAN Xiaoyan, LI Mingzhe, CHEN Long, LIANG Chao, LU Wenli, PAN Zhiqiang, JIA Dongwei, PENG Peike, FANG Zhaoqin, LIU Xiaomei. Transcriptome profile and its partly verification of human hepatocellular carcinoma cells exposed to Yuzhizi (Fructus Akebiae Quinatae) seed extract[J]. Journal of Traditional Chinese Medicine, 2022, 42(6): 922-931.
share this article
| Gene | Primer sequence |
|---|---|
| Ribosomal protein S27a (RPS27A) | Sense 5’-GCCCTTCTGATGAATGTGGT-3’ |
| Anti-sense 5’-TGGTGTGACCATCAATTTGG-3’ | |
| Transferrin (TF) | Sense 5’-GCTGCTTTGGGAGACAGAAC-3’ |
| Anti-sense 5’-ACAGCTTAAGATCGGCCTCA-3’ | |
| Ribosomal protein S20 (RPS20) | Sense 5’-AAGGGCTGAGGATTTTTGGT-3’ |
| Anti-sense 5’-GGGTGTTTTTCCGGTATCCT-3’ | |
| Ribosomal protein L9 (RPL9) | Sense 5’-ATGCTCACTTCCCCATCAAC-3’ |
| Anti-sense 5’-ACACCTGGTCTCATCCGAAC-3’ | |
| Protein phosphatase 2 regulatory Subunit b alpha (PPP2R2A) | Sense 5’-AGTCAGTTTGGGGTCAGTGG-3’ |
| Anti-sense 5’-TGCCCAAAGGTATGACACAA-3’ | |
| Transthyretin (TTR) | Sense 5’-ATGGCTTCTCATCGTCTGCT-3’ |
| Anti-sense 5’-TGTCATCAGCAGCCTTTCTG-3’ | |
| Thioredoxin reductase 1 (TXNRD1) | Sense 5’-AGGAAGATTGCTGGCTCAGA-3’ |
| Anti-sense 5’-CTCCTCAGAAAGGCCACAAG-3’ | |
| Ribosomal protein L3 (RPL3) | Sense 5’-CCGCACTGAGATCAACAAGA-3’ |
| Anti-sense 5’-CAGCCTTTCAGCATGACAAA-3’ | |
| Ribophorin I (RPN1) | Sense 5’-GCACATTCAGGACATTGTGG-3’ |
| Anti-sense 5’-CCACTGTTGAGGGTGGAGAT-3’ | |
| Ribosomal protein L24 (RPL24) | Sense 5’-CAAATTCCAGAGGGCCATTA-3’ |
| Anti-sense 5’-GCAGCCCTGATAGCTTGTTC-3’ | |
| Alpha fetoprotein (AFP) | Sense 5’-GTAGCGCTGCAAACAATGAA-3’ |
| Anti-sense 5’-CCCTCTTCAGCAAAGCAGAC-3’ | |
| Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) | Sense 5’-CGACCACTTTGTCAAGCTCA-3’ |
| Anti-sense 5’-AGGGGTCTACATGGCAACTG-3’ |
Table 1 Primers used in polymerase chain reaction
| Gene | Primer sequence |
|---|---|
| Ribosomal protein S27a (RPS27A) | Sense 5’-GCCCTTCTGATGAATGTGGT-3’ |
| Anti-sense 5’-TGGTGTGACCATCAATTTGG-3’ | |
| Transferrin (TF) | Sense 5’-GCTGCTTTGGGAGACAGAAC-3’ |
| Anti-sense 5’-ACAGCTTAAGATCGGCCTCA-3’ | |
| Ribosomal protein S20 (RPS20) | Sense 5’-AAGGGCTGAGGATTTTTGGT-3’ |
| Anti-sense 5’-GGGTGTTTTTCCGGTATCCT-3’ | |
| Ribosomal protein L9 (RPL9) | Sense 5’-ATGCTCACTTCCCCATCAAC-3’ |
| Anti-sense 5’-ACACCTGGTCTCATCCGAAC-3’ | |
| Protein phosphatase 2 regulatory Subunit b alpha (PPP2R2A) | Sense 5’-AGTCAGTTTGGGGTCAGTGG-3’ |
| Anti-sense 5’-TGCCCAAAGGTATGACACAA-3’ | |
| Transthyretin (TTR) | Sense 5’-ATGGCTTCTCATCGTCTGCT-3’ |
| Anti-sense 5’-TGTCATCAGCAGCCTTTCTG-3’ | |
| Thioredoxin reductase 1 (TXNRD1) | Sense 5’-AGGAAGATTGCTGGCTCAGA-3’ |
| Anti-sense 5’-CTCCTCAGAAAGGCCACAAG-3’ | |
| Ribosomal protein L3 (RPL3) | Sense 5’-CCGCACTGAGATCAACAAGA-3’ |
| Anti-sense 5’-CAGCCTTTCAGCATGACAAA-3’ | |
| Ribophorin I (RPN1) | Sense 5’-GCACATTCAGGACATTGTGG-3’ |
| Anti-sense 5’-CCACTGTTGAGGGTGGAGAT-3’ | |
| Ribosomal protein L24 (RPL24) | Sense 5’-CAAATTCCAGAGGGCCATTA-3’ |
| Anti-sense 5’-GCAGCCCTGATAGCTTGTTC-3’ | |
| Alpha fetoprotein (AFP) | Sense 5’-GTAGCGCTGCAAACAATGAA-3’ |
| Anti-sense 5’-CCCTCTTCAGCAAAGCAGAC-3’ | |
| Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) | Sense 5’-CGACCACTTTGTCAAGCTCA-3’ |
| Anti-sense 5’-AGGGGTCTACATGGCAACTG-3’ |
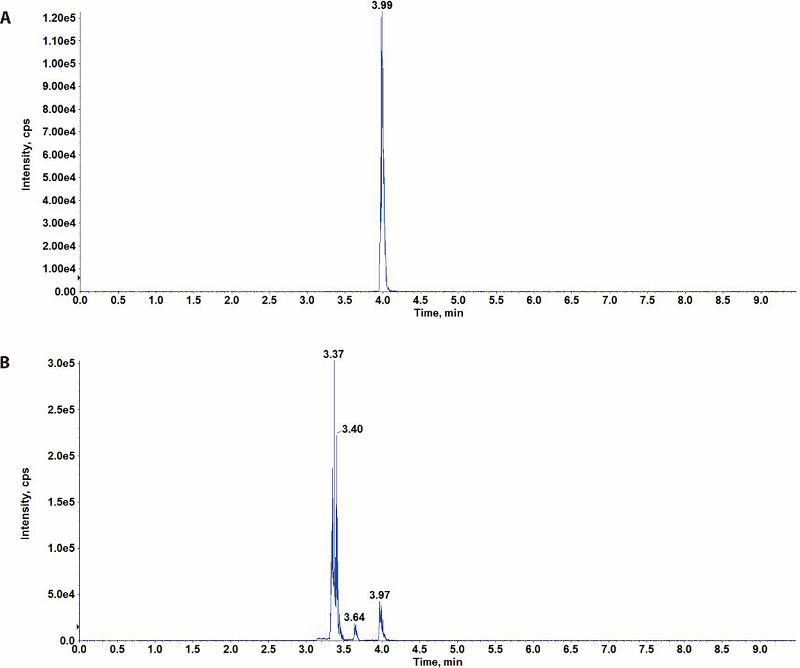
Figure 1 Chromatography of α-hederin standard and FAQSE sample by UPLC-ESI-MS/MS A: α-hederin standard solution, the injection concentration of which was 25, 50, 100, 200 and 400 ng/mL; B: FAQSE sample solution, the injection concentration of which was 5.0 mg/mL; FAQSE: Yuzhizi (Fructus Akebiae Quinatae) seed extract; UPLC-ESI-MS/MS: ultra-performance liquid chromatography-electrospray ionization tandem mass spectrometry.
| Group | n | HepG2 | Huh7 | |||||
|---|---|---|---|---|---|---|---|---|
| 24 h | 48 h | 24 h | 48 h | |||||
| Control | 3 | 100.0±4.6 | 100.0±2.0 | 100.0±1.6 | 100.0±1.3 | |||
| FAQSE | 0.3125 mg/mL | 3 | 97.8±3.4 | 93.0±1.9 | 97.1±1.2 | 86.2±4.6 | ||
| 0.625 mg/mL | 3 | 82.0±4.1a | 61.4±6.0a | 70.1±3.1a | 50.4±1.4a | |||
| 0.9375 mg/mL | 3 | 70.5±2.3a | 51.6±1.9a | 61.1±4.9a | 40.2±1.2a | |||
| 1.25 mg/mL | 3 | 57.6±3.6a | 40.3±0.7a | 48.6±2.1a | 30.8±1.0a | |||
Table 2 Effect of FAQSE on the viability of HepG2 and Huh7 cells (%, $\bar{x} \pm s$)
| Group | n | HepG2 | Huh7 | |||||
|---|---|---|---|---|---|---|---|---|
| 24 h | 48 h | 24 h | 48 h | |||||
| Control | 3 | 100.0±4.6 | 100.0±2.0 | 100.0±1.6 | 100.0±1.3 | |||
| FAQSE | 0.3125 mg/mL | 3 | 97.8±3.4 | 93.0±1.9 | 97.1±1.2 | 86.2±4.6 | ||
| 0.625 mg/mL | 3 | 82.0±4.1a | 61.4±6.0a | 70.1±3.1a | 50.4±1.4a | |||
| 0.9375 mg/mL | 3 | 70.5±2.3a | 51.6±1.9a | 61.1±4.9a | 40.2±1.2a | |||
| 1.25 mg/mL | 3 | 57.6±3.6a | 40.3±0.7a | 48.6±2.1a | 30.8±1.0a | |||
| Pathway ID | Name | Count | P value | Gene |
|---|---|---|---|---|
| hsa04144 | Endocytosis | 10 | 0.002997 | GIT1, RAB7A, RAB31, RAB5B, ARRB2, ARF3, EPS15L1, STAM, HSPA1A, CAPZB |
| hsa00480 | Glutathione metabolism | 3 | 0.133177 | GSTM4, G6PD, MGST1 |
| hsa04141 | Protein processing in endoplasmic reticulum | 5 | 0.160565 | UBQLN4, HSPA1A, DNAJB1, CAPN2, PPP1R15A |
| hsa04721 | Synaptic vesicle cycle | 3 | 0.1859 | STX3, STXBP1, ATP6V0D1 |
| hsa04142 | Lysosome | 4 | 0.192439 | CD68, IDS, ATP6V0D1, GGA3 |
Table 3 Top 5 KEGG pathways of co-upregulated DEGs
| Pathway ID | Name | Count | P value | Gene |
|---|---|---|---|---|
| hsa04144 | Endocytosis | 10 | 0.002997 | GIT1, RAB7A, RAB31, RAB5B, ARRB2, ARF3, EPS15L1, STAM, HSPA1A, CAPZB |
| hsa00480 | Glutathione metabolism | 3 | 0.133177 | GSTM4, G6PD, MGST1 |
| hsa04141 | Protein processing in endoplasmic reticulum | 5 | 0.160565 | UBQLN4, HSPA1A, DNAJB1, CAPN2, PPP1R15A |
| hsa04721 | Synaptic vesicle cycle | 3 | 0.1859 | STX3, STXBP1, ATP6V0D1 |
| hsa04142 | Lysosome | 4 | 0.192439 | CD68, IDS, ATP6V0D1, GGA3 |
| Pathway ID | Name | Count | P value | Gene |
|---|---|---|---|---|
| hsa03010 | Ribosome | 11 | 1.16×10-8 | RPSA, RPL17, RPS27, RPS17, RPL9, RPL3, RPL24, RPS20, RPS21, RPS27A, RPS8 |
| hsa01230 | Biosynthesis of amino acids | 4 | 0.011393 | PYCR1, ALDH18A1, ASS1, ALDOC |
| hsa00330 | Arginine and proline metabolism | 3 | 0.042035 | PYCR1, ALDH18A1, AGMAT |
| hsa05322 | Systemic lupus erythematosus | 4 | 0.056687 | HIST1H2BM, C5, HIST1H3B, HIST1H4C |
| hsa04610 | Complement and coagulation cascades | 3 | 0.074505 | FGG, FGA, C5 |
Table 4 Top 5 KEGG pathways of co-downregulated DEGs
| Pathway ID | Name | Count | P value | Gene |
|---|---|---|---|---|
| hsa03010 | Ribosome | 11 | 1.16×10-8 | RPSA, RPL17, RPS27, RPS17, RPL9, RPL3, RPL24, RPS20, RPS21, RPS27A, RPS8 |
| hsa01230 | Biosynthesis of amino acids | 4 | 0.011393 | PYCR1, ALDH18A1, ASS1, ALDOC |
| hsa00330 | Arginine and proline metabolism | 3 | 0.042035 | PYCR1, ALDH18A1, AGMAT |
| hsa05322 | Systemic lupus erythematosus | 4 | 0.056687 | HIST1H2BM, C5, HIST1H3B, HIST1H4C |
| hsa04610 | Complement and coagulation cascades | 3 | 0.074505 | FGG, FGA, C5 |

Figure 2 PPI network of some important co-DEGs Genes with pink and blue background were upregulated and downregulated genes, respectively. PPI: protein-protein interaction; DEGs: differentially expressed genes.
| Gene | HepG2 | Huh7 | ||||
|---|---|---|---|---|---|---|
| Control group (n = 3) | FAQSE group (n = 3) | Control group (n = 3) | FAQSE group (n = 3) | |||
| RPS27A | 1.00±0.12 | 0.64±0.03a | 1.00±0.12 | 0.81±0.18 | ||
| TF | 1.00±0.17 | 0.21±0.02a | 1.00±0.10 | 0.38±0.18a | ||
| RPS20 | 1.00±0.36 | 0.73±0.10 | 1.00±0.00 | 0.96±0.16 | ||
| RPL9 | 1.00±0.25 | 0.75±0.14 | 1.00±0.13 | 0.93±0.10 | ||
| PPP2R2A | 1.00±0.18 | 0.97±0.06 | 1.00±0.26 | 0.76±0.17 | ||
| TTR | 1.00±0.28 | 0.45±0.04 | 1.00±0.69 | 0.27±0.03 | ||
| TXNRD1 | 1.00±0.14 | 1.56±0.24a | 1.00±0.15 | 2.04±0.53a | ||
| RPL3 | 1.00±0.33 | 0.58±0.11a | 1.00±0.00 | 1.05±0.30 | ||
| RPN1 | 1.00±0.15 | 0.69±0.12 | 1.00±0.07 | 1.29±0.26 | ||
| RPL24 | 1.00±0.42 | 0.68±0.05 | 1.00±0.07 | 1.46±0.46 | ||
| AFP | 1.00±0.26 | 0.32±0.03a | 1.00±0.07 | 0.22±0.04a | ||
Table 5 mRNA expression of some important co-DEGs in HepG2 and Huh7 cells (relative expression compared to control group, $\bar{x} \pm s$)
| Gene | HepG2 | Huh7 | ||||
|---|---|---|---|---|---|---|
| Control group (n = 3) | FAQSE group (n = 3) | Control group (n = 3) | FAQSE group (n = 3) | |||
| RPS27A | 1.00±0.12 | 0.64±0.03a | 1.00±0.12 | 0.81±0.18 | ||
| TF | 1.00±0.17 | 0.21±0.02a | 1.00±0.10 | 0.38±0.18a | ||
| RPS20 | 1.00±0.36 | 0.73±0.10 | 1.00±0.00 | 0.96±0.16 | ||
| RPL9 | 1.00±0.25 | 0.75±0.14 | 1.00±0.13 | 0.93±0.10 | ||
| PPP2R2A | 1.00±0.18 | 0.97±0.06 | 1.00±0.26 | 0.76±0.17 | ||
| TTR | 1.00±0.28 | 0.45±0.04 | 1.00±0.69 | 0.27±0.03 | ||
| TXNRD1 | 1.00±0.14 | 1.56±0.24a | 1.00±0.15 | 2.04±0.53a | ||
| RPL3 | 1.00±0.33 | 0.58±0.11a | 1.00±0.00 | 1.05±0.30 | ||
| RPN1 | 1.00±0.15 | 0.69±0.12 | 1.00±0.07 | 1.29±0.26 | ||
| RPL24 | 1.00±0.42 | 0.68±0.05 | 1.00±0.07 | 1.46±0.46 | ||
| AFP | 1.00±0.26 | 0.32±0.03a | 1.00±0.07 | 0.22±0.04a | ||
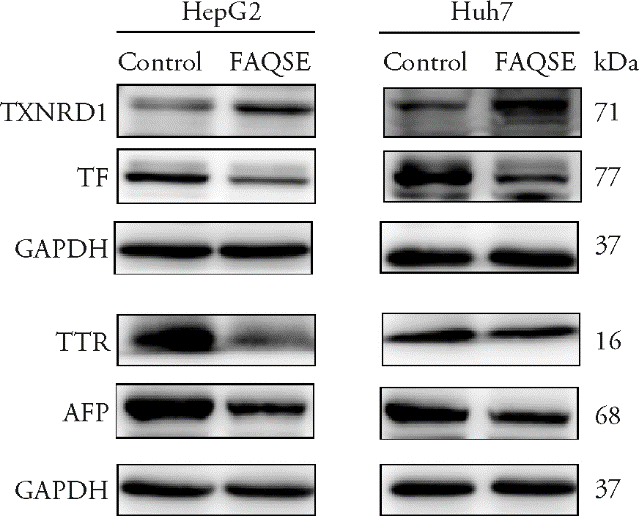
Figure 3 Protein expression of some important co-DEGs in HepG2 and Huh7 cells Control group: treated only with normal medium; FAQSE group: treated with FAQSE 0.625 mg/mL for 48 h; DEGs: differentially expressed genes; FAQSE: Yuzhizi (Fructus Akebiae Quinatae) seed extract; TXNRD1: thioredoxin reductase 1; TF: transferrin; TTR: transthyretin; AFP: alpha fetoprotein; GAPDH: glyceraldehyde-3-phosphate dehydrogenase.
| Protein | HepG2 | Huh7 | |||
|---|---|---|---|---|---|
| Control group (n = 3) | FAQSE group (n = 3) | Control group (n = 3) | FAQSE group (n = 3) | ||
| TXNRD1 | 1.10±0.75 | 1.37±0.52 | 0.91±0.32 | 1.14±0.16 | |
| TTR | 1.08±0.08 | 0.68±0.21a | 0.65±0.16 | 0.76±0.17 | |
| TF | 1.17±0.87 | 0.70±0.15 | 1.11±0.42 | 0.59±0.32 | |
| AFP | 1.12±0.37 | 0.93±0.16 | 1.21±0.48 | 0.53±0.26 | |
Table 6 Relative protein expression of some important co-DEGs in HepG2 and Huh7 cells (protein/GAPDH, $\bar{x} \pm s$)
| Protein | HepG2 | Huh7 | |||
|---|---|---|---|---|---|
| Control group (n = 3) | FAQSE group (n = 3) | Control group (n = 3) | FAQSE group (n = 3) | ||
| TXNRD1 | 1.10±0.75 | 1.37±0.52 | 0.91±0.32 | 1.14±0.16 | |
| TTR | 1.08±0.08 | 0.68±0.21a | 0.65±0.16 | 0.76±0.17 | |
| TF | 1.17±0.87 | 0.70±0.15 | 1.11±0.42 | 0.59±0.32 | |
| AFP | 1.12±0.37 | 0.93±0.16 | 1.21±0.48 | 0.53±0.26 | |
| Group | n | HepG2 | Huh7 |
|---|---|---|---|
| Control | 3 | 284±7 | 95±3 |
| FAQSE | 3 | 242±17a | 64±5a |
Table 7 FAQSE reduced the AFP release of HepG2 and Huh7 cells
| Group | n | HepG2 | Huh7 |
|---|---|---|---|
| Control | 3 | 284±7 | 95±3 |
| FAQSE | 3 | 242±17a | 64±5a |
| [1] |
Sung H, Ferlay J, Siegel RL, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA-Cancer J Clin 2021; 71: 209-49.
DOI URL |
| [2] |
Ting CT, Li WC, Chen CY, Tsai TH. Preventive and therapeutic role of traditional Chinese herbal medicine in hepatocellular carcinoma. J Chin Med Assoc 2015; 78: 139-44.
DOI PMID |
| [3] |
Wu P, Dugoua JJ, Eyawo O, Mills EJ. Traditional Chinese Medicines in the treatment of hepatocellular cancers: a systematic review and Meta-analysis. J Exp Clin Cancer Res 2009; 28: 112.
DOI URL |
| [4] | National Pharmacopoeia Commission. Pharmacopoeia of the People's Republic of China (Part I). Beijing: China Medical Science and Technology Press, 2015: 297 |
| [5] | Lu WL, Ren HY, Liang C, et al. Akebia trifoliate (Thunb.) Koidz seed extract inhibits the proliferation of human hepatocellular carcinoma cell lines via inducing endoplasmic reticulum stress. Evid Based Complement Alternat Med 2014; 2014: 192749. |
| [6] |
Lu WL, Yang T, Song QJ, et al. Akebia trifoliata (Thunb.) Koidz seed extract inhibits human hepatocellular carcinoma cell migration and invasion in vitro. J Ethnopharmacol 2019; 234: 204-15.
DOI URL |
| [7] | Oliveros JC. Venny. An interactive tool for comparing lists with Venn's diagrams. 2007-2015, cited 2021-07-15; Available from URL: https://bioinfogp.cnb.csic.es/tools/venny/index.html. |
| [8] |
Huang DW, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 2009; 4: 44-57.
DOI PMID |
| [9] |
Szklarczyk D, Gable AL, Lyon D, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 2019; 47: D607-13.
DOI |
| [10] |
Kohl M, Wiese S, Warscheid B. Cytoscape: software for visualization and analysis of biological networks. Methods Mol Biol 2011; 696: 291-303.
DOI PMID |
| [11] |
Smart JA, Oleksak JE, Hartsough EJ. Cell adhesion molecules in plasticity and metastasis. Mol Cancer Res 2021; 19: 25-37.
DOI PMID |
| [12] |
Cassiman D, van Pelt J, De Vos R, et al. Synaptophysin: A novel marker for human and rat hepatic stellate cells. Am J Pathol 1999; 155: 1831-9.
PMID |
| [13] |
Kennedy L, Sandhu JK, Harper ME, Cuperlovic-Culf M. Role of glutathione in cancer: from mechanisms to therapies. Biomolecules 2020; 10: 1429.
DOI URL |
| [14] |
Ding Z, Ericksen RE, Escande-Beillard N, et al. Metabolic pathway analyses identify proline biosynthesis pathway as a promoter of liver tumorigenesis. J Hepatol 2020; 72: 725-35.
DOI PMID |
| [15] |
Pio R, Corrales L, Lambris JD. The role of complement in tumor growth. Adv Exp Med Biol 2014; 772: 229-62.
DOI PMID |
| [16] |
Woo SR, Corrales L, Gajewski TF. Innate immune recognition of cancer. Annu Rev Immunol 2015; 33: 445-74.
DOI URL |
| [17] |
Ruggero D, Pandolfi PP. Does the ribosome translate cancer?. Nat Rev Cancer 2003; 3: 179-92.
DOI PMID |
| [18] |
Cordon CC, Reuter VE. Alterations of tumor suppressor genes in bladder cancer. Semin Diagn Pathol 1997; 14: 123-32.
PMID |
| [19] |
Kuo YC, Huang KY, Yang CH, Yang YS, Lee WY, Chiang CW. Regulation of phosphorylation of Thr-308 of Akt, cell proliferation, and survival by the B55alpha regulatory subunit targeting of the protein phosphatase 2A holoenzyme to Akt. J Biol Chem 2008; 283: 1882-92.
DOI URL |
| [20] |
Wong QW, Ching AK, Chan AW, et al. MiR-222 overexpression confers cell migratory advantages in hepatocellular carcinoma through enhancing AKT signaling. Clin Cancer Res 2010; 16: 867-75.
DOI PMID |
| [21] |
Kim J, Bae JS. ROS homeostasis and metabolism: a critical liaison for cancer therapy. Exp Mol Med 2016; 48: e269.
DOI URL |
| [22] |
Park S, Yoon SY, Kim KE, et al. Interleukin-18 induces transferrin expression in breast cancer cell line MCF-7. Cancer Lett 2009; 286: 189-95.
DOI PMID |
| [23] |
Qiao W, Leng F, Liu T, et al. Prognostic value of prealbumin in liver cancer: a systematic review and meta-analysis. Nutr Cancer 2020; 72: 909-16.
DOI PMID |
| [24] |
Bai DS, Zhang C, Chen P, Jin SJ, Jiang GQ. The prognostic correlation of AFP level at diagnosis with pathological grade, progression, and survival of patients with hepatocellular carcinoma. Sci Rep 2017; 7: 12870.
DOI URL |
| [25] |
Zheng Y, Zhu M, Li M. Effects of alpha-fetoprotein on the occurrence and progression of hepatocellular carcinoma. J Cancer Res Clin Oncol 2020; 146: 2439-46.
DOI PMID |
| [26] | Kanda S, Shiroi A, Ouji Y, et al. In vitro differentiation of hepatocyte-like cells from embryonic stem cells promoted by gene transfer of hepatocyte nuclear factor 3β. Hepatol Res 2003; 26: 225-31. |
| [27] |
Uriel J, Iskandar Y, Laborda J, et al. Defective uptake of alpha-fetoprotein (AFP) and transferrin (Tf) by PHA-activated peripheral blood lymphocytes from patients with AIDS and related syndromes. AIDS Res Hum Retrov 1990; 6: 401-10.
PMID |
| [28] |
Adachi M, Kai K, Yamaji K, et al. Transferrin receptor 1 overexpression is associated with tumour de-differentiation and acts as a potential prognostic indicator of hepatocellular carcinoma. Histopathology 2019; 75: 63-73.
DOI PMID |
| [1] | QU Dongxiao, GE Yiqin, ZHUO Limin, CHEN Liji, XUE Yonghua, CHENG Jiwei, TAO Jie, LI Guoyi, ZHU Yudan, XIAO Qian. Efficacy and mechanisms of Dingxian pill (定痫丸) combined with valproic acid on pentylenetetrazol-induced chronic epilepsy in rats [J]. Journal of Traditional Chinese Medicine, 2023, 43(2): 286-294. |
| [2] | ZHANG Jiri Mutu, LIANG Shilong, NIE Peng, LIAO Yong’an, AI Qinying, YAN Xiaojun, LIU Hongning, JI Yanhua, ZENG Zhijun. Efficacy of Kushen decoction (苦参汤) on high-fat-diet-induced hyperlipidemia in rats [J]. Journal of Traditional Chinese Medicine, 2022, 42(3): 364-371. |
| [3] | ZHANG Chengfei, QIN Lingling, WANG Haiyan, SUN Boju, ZHAO Dan, ZHANG Qiue, ZHONG Fengying, WU Lili, LIU Tonghua. Efficacy of aqueous extract of flower of Edgeworthia gardneri (Wall.) Meisn on glucose and lipid metabolism in KK/Upj-Ay/J mice [J]. Journal of Traditional Chinese Medicine, 2022, 42(2): 187-193. |
| [4] | Xiaomei Liu, Zhaoqin Fang, Zhiqiang Pan, Wenli Lu, Zhonghua Wu, Chao Liang, Yuanyuan Zhang. Pituitary transcriptome profile of liver cancer mice with different syndromes reveals the relevance of pituitary to the cancer and syndromes [J]. Journal of Traditional Chinese Medicine, 2014, 34(06): 691-698. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||

