Journal of Traditional Chinese Medicine ›› 2024, Vol. 44 ›› Issue (2): 277-288.DOI: 10.19852/j.cnki.jtcm.20231018.001
Previous Articles Next Articles
Gehua Jiejiu Dizhi decoction (葛花解酒涤脂汤) ameliorates alcoholic fatty liver in mice by regulating lipid and bile acid metabolism and with exertion of antioxidant stress based on 4DLabel-free quantitative proteomic study
HAN Min1, YI Xu2( ), YOU Shaowei2, WU Xueli2, WANG Shuoshi2, HE Diancheng2
), YOU Shaowei2, WU Xueli2, WANG Shuoshi2, HE Diancheng2
- 1 Guizhou University of Traditional Chinese Medicine, Graduate School, Guiyang 550025, China
2 Department of Clinical medical laboratory, Department of Gastroenterology, the Second Affiliated Hospital of Guizhou University of Traditional Chinese Medicine, Guiyang 550003, China
-
Received:2022-11-11Accepted:2023-04-27Online:2024-04-15Published:2023-12-18 -
Contact:Prof. YI Xu, Department of Clinical medical laboratory, the Second Affiliated Hospital of Guizhou University of Traditional Chinese Medicine, Guiyang 550003, China.yixu2013@yeah.net Telephone: +86-851-85512704 -
Supported by:National Science Foundation-funded Project: the Study on the Changes of Energy Metabolism and Molecular Regulation Mechanism of Alcoholic Fatty Liver based on Sirtuins1-Adenosine Monophosphate-Activated Protein Kinase Signal System and the Intervention of Gehua Jiejiu dizhi decoction(81660752);Basic Research Project of Guizhou Provincial Science and Technology Plan: Study on the Mechanism of Sirtuins1 Mediated Deacetylation in the Regulation of Alcoholic Fatty Liver Metabolism and the Intervention of Gehua Jiejiu Dizhi Tang(QianKeHe Fundamentals-ZK [2023] General 410)
Cite this article
HAN Min, YI Xu, YOU Shaowei, WU Xueli, WANG Shuoshi, HE Diancheng. Gehua Jiejiu Dizhi decoction (葛花解酒涤脂汤) ameliorates alcoholic fatty liver in mice by regulating lipid and bile acid metabolism and with exertion of antioxidant stress based on 4DLabel-free quantitative proteomic study[J]. Journal of Traditional Chinese Medicine, 2024, 44(2): 277-288.
share this article
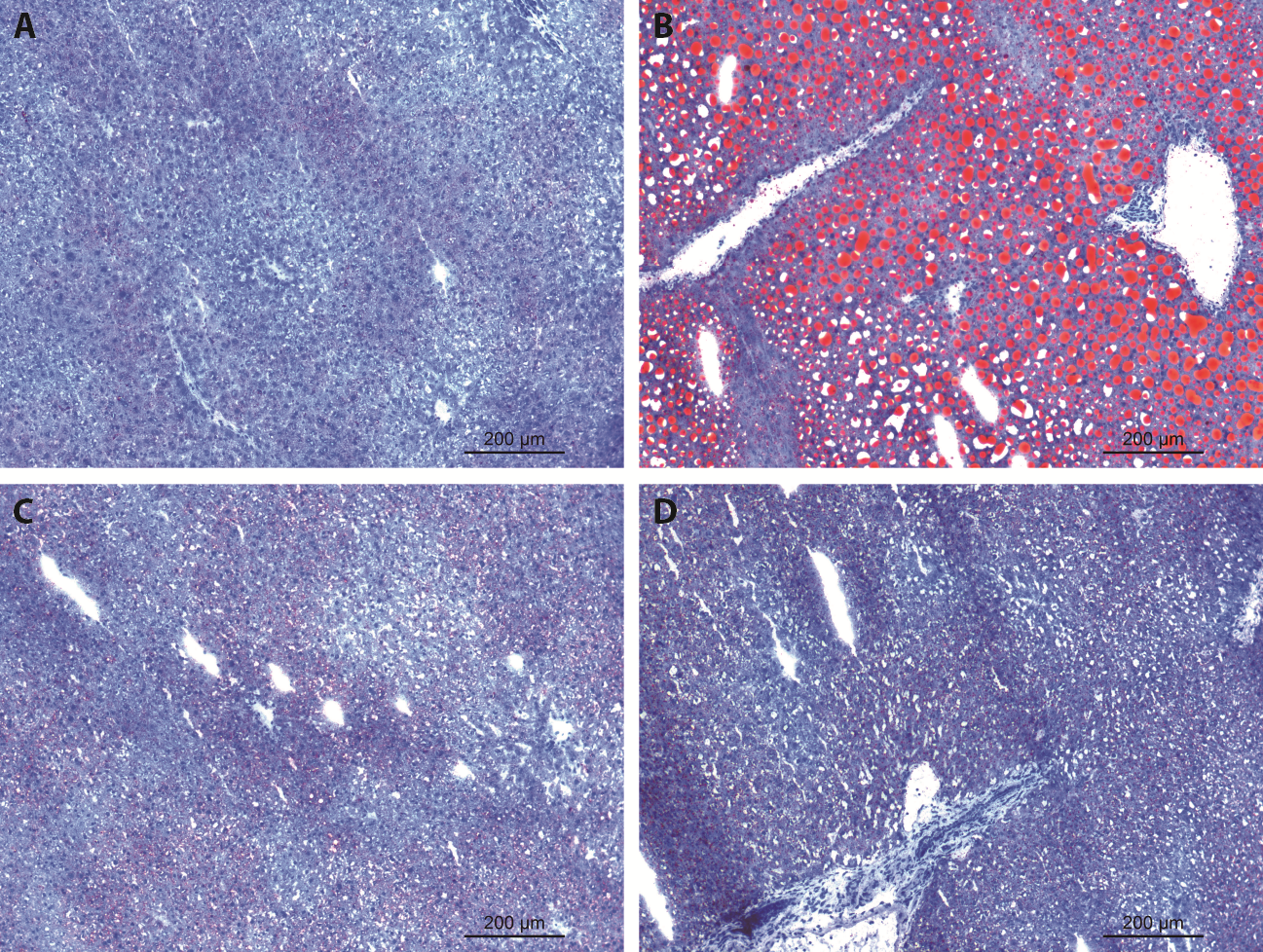
Figure 1 Hepatic steatosis of liver in 4 groups of mice by ORO staining (× 100 magnification) A: control group:treated only withfree drinking water; B: AFLD model group:treated with ethanol and drinking water (2.6 g/kg); C: GJDD intervention group: treated with GJDD of 4.9 g/kg; D: resveratrol intervention group: treated with resveratrol of 4 g/kg. Diffuse and red dropwise lipid vacuoles were found in the AFLD mice liver tissue, but they were significantly reduced both in GJDD and resveratrol-treated mice, accompanied by small drops, especially in the GJDD group. Semiquantitative analysis of fatty degeneration showed that AFLD mice were mainly MiS, and the moderate MaS was observed also in 32.2% of mice. Correspondingly, GJDD group and resveratrol group showed a small amount of MiS and no MaS. Black arrow indicates large steatosis in model group, but not in other groups (× 200 magnification). ORO: oil red o; MaS: macrovesicular steatosis; MiS: microvesicular steatosis.
| Group | n | MaS (%) | MiS (%) | ToS (%) | ||
|---|---|---|---|---|---|---|
| Mild | Moderate | Severe | ||||
| Control | 8 | 0 | 0 | 0 | 2.0±1.2 | 2.0±1.2 |
| Model | 8 | 10.0±2.5a | 32.4±6.5a | 0 | 36.3±7.6a | 68.8±10.4a |
| GJDD | 8 | 0.9±1.2 | 0.3±6.0 | 0 | 5.0±2.6b | 7.2±2.7b |
| Resveratrol | 8 | 0.9±2.3 | 0.3±4.0 | 0 | 10.3±2.7b | 18.6±3.2b |
Table 1 Results of semiquantitative analysis of liver steatosis by ORO staining in each group ( ± s)
| Group | n | MaS (%) | MiS (%) | ToS (%) | ||
|---|---|---|---|---|---|---|
| Mild | Moderate | Severe | ||||
| Control | 8 | 0 | 0 | 0 | 2.0±1.2 | 2.0±1.2 |
| Model | 8 | 10.0±2.5a | 32.4±6.5a | 0 | 36.3±7.6a | 68.8±10.4a |
| GJDD | 8 | 0.9±1.2 | 0.3±6.0 | 0 | 5.0±2.6b | 7.2±2.7b |
| Resveratrol | 8 | 0.9±2.3 | 0.3±4.0 | 0 | 10.3±2.7b | 18.6±3.2b |
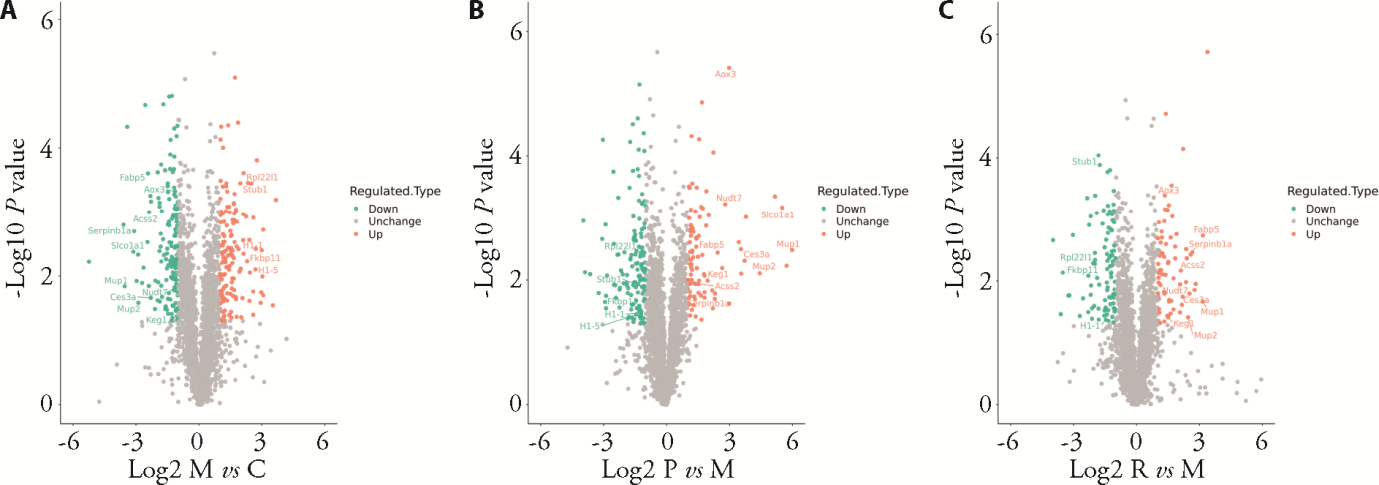
Figure 2 Volcano plots showed the differentially protein expression in every two groups (M/C, P/M, R/M) A: AFLD group/Control group (M/C); B: GJDD group/AFLD group (P/M); C: resveratrol group/AFLD group (R/M). Control group: treated only with free drinking water; AFLD model group: treated with ethanol and drinking water (2.6 g/kg); GJDD intervention group: treated with GJDD of 4.9 g/kg; resveratrol intervention group: treated with resveratrol of 4 g/kg. AFLD: alcoholic fatty live disease; GJDD: Gehua Jiejiu Dizhi decoction. The ratio of the mean value of repeated quantitative value of protein in the comparison sample pair was taken as the fold change. The relative quantitative value of protein in the comparison sample pair was tested by t-test. The specified thresholds include log2 fold change and adjusted P-value after log conversion, which were used to measure the fold and significance of protein expression differences respectively. In the figure, positive and red dots indicated that the difference was up-regulated. On the contrary, negative and green dots indicated that the difference was down-regulated.

Figure 3 Bubble plot of KEGG pathway enrichment analysis of proteins with differential expression level A: AFLD group/Control group; B: GJDD group/AFLD group; C: resveratrol group/AFLD group. Control group: treated only with free drinking water; AFLD model group: treated with ethanol and drinking water (2.6 g/kg); GJDD intervention group: treated with GJDD of 4.9 g/kg; resveratrol intervention group: treated with resveratrol of 4 g/kg. X-axis represents enrichment factor. Y-axis represents pathway name. The darker the mapping color, the smaller the P-value and the more obvious the significance, the larger the mapping value and the more enrichment. The same KEGG enrichment pathway involved in the differential proteins of M/C, P/M, and R/M comparison groups are boxed in three colors respectively, in which red is “retinol metabolism”, purple is “pyruvate metabolism”, and blue is “steroid hormone biosynthesis”. KEGG: Kyoto Encyclopedia of Genes and Genomes; AFLD: alcoholic fatty live disease; GJDD: Gehua Jiejiu Dizhi decoction.
| Proteins accession | Protein description/ Gene | MW (kDa) | Coverage (%) | Unique peptides | Ratio | ||
|---|---|---|---|---|---|---|---|
| M/C | P/C | P/M | |||||
| P43275 | Histone H1.1/ H1-1 | 21.79 | 31.9 | 4 | 6.54 | 2.81 | 0.43 |
| P43276 | Histone H1.5/ H1-5 | 22.58 | 25.1 | 4 | 6.48 | 2.37 | 0.37 |
| Q9WUD1 | STIP1 homology and U box-containing protein/ Stub1 | 34.91 | 34.2 | 9 | 5.66 | 2.55 | 0.45 |
| Q9D7S7 | 60S ribosomal protein L22-like 1/ Rpl22l1 | 14.47 | 45.1 | 4 | 5.08 | 2.30 | 0.45 |
| Q9D1M7 | Peptidyl-prolyl cis-trans isomerase/ Fkbp11 | 22.14 | 41.3 | 5 | 4.91 | 1.85 | 0.38 |
| G3X982 | Aldehyde oxidase 3/ Aox3 | 146.9 | 55.4 | 52 | 0.46 | 3.60 | 7.9 |
| Q99P30 | Peroxisomal coenzyme A/ Nudt7 | 26.86 | 51.7 | 14 | 0.45 | 3.11 | 6.9 |
| Q9DCY0 | Glycine N-acyltransfera se-like protein/ Keg1 | 33.72 | 55.3 | 13 | 0.40 | 2.53 | 6.28 |
| Q63880 | Carboxylesterase 3A/Ces3a | 63.32 | 52.7 | 11 | 0.20 | 2.68 | 13.17 |
| Q05816 | Fattyacid-bindingprotein 5/ Fabp5 | 15.14 | 72.6 | 9 | 0.19 | 0.50 | 2.66 |
| Q9QXG4 | Acetyl-coenzyme A synthe tase/ Acss2 | 78.86 | 43.2 | 25 | 0.19 | 0.48 | 2.48 |
| P11589 | Major urinary protein 2/ Mup2 | 20.66 | 78.3 | 3 | 0.14 | 2.94 | 21.74 |
| Q9QXZ6 | Solute carrier organican ion transporter family member/ Slco1a1 | 74.40 | 32.1 | 16 | 0.11 | 5.21 | 45.54 |
| P11588 | Major urinary protein 1/ Mup1 | 20.65 | 81.1 | 3 | 0.09 | 5.46 | 63.4 |
| Q9D154 | Leukocyte elastase inhibitor A/ Serpinb1a | 42.58 | 35.4 | 12 | 0.08 | 0.26 | 3.11 |
Table 2 Differential co-expressed proteins in groups of C, M and P
| Proteins accession | Protein description/ Gene | MW (kDa) | Coverage (%) | Unique peptides | Ratio | ||
|---|---|---|---|---|---|---|---|
| M/C | P/C | P/M | |||||
| P43275 | Histone H1.1/ H1-1 | 21.79 | 31.9 | 4 | 6.54 | 2.81 | 0.43 |
| P43276 | Histone H1.5/ H1-5 | 22.58 | 25.1 | 4 | 6.48 | 2.37 | 0.37 |
| Q9WUD1 | STIP1 homology and U box-containing protein/ Stub1 | 34.91 | 34.2 | 9 | 5.66 | 2.55 | 0.45 |
| Q9D7S7 | 60S ribosomal protein L22-like 1/ Rpl22l1 | 14.47 | 45.1 | 4 | 5.08 | 2.30 | 0.45 |
| Q9D1M7 | Peptidyl-prolyl cis-trans isomerase/ Fkbp11 | 22.14 | 41.3 | 5 | 4.91 | 1.85 | 0.38 |
| G3X982 | Aldehyde oxidase 3/ Aox3 | 146.9 | 55.4 | 52 | 0.46 | 3.60 | 7.9 |
| Q99P30 | Peroxisomal coenzyme A/ Nudt7 | 26.86 | 51.7 | 14 | 0.45 | 3.11 | 6.9 |
| Q9DCY0 | Glycine N-acyltransfera se-like protein/ Keg1 | 33.72 | 55.3 | 13 | 0.40 | 2.53 | 6.28 |
| Q63880 | Carboxylesterase 3A/Ces3a | 63.32 | 52.7 | 11 | 0.20 | 2.68 | 13.17 |
| Q05816 | Fattyacid-bindingprotein 5/ Fabp5 | 15.14 | 72.6 | 9 | 0.19 | 0.50 | 2.66 |
| Q9QXG4 | Acetyl-coenzyme A synthe tase/ Acss2 | 78.86 | 43.2 | 25 | 0.19 | 0.48 | 2.48 |
| P11589 | Major urinary protein 2/ Mup2 | 20.66 | 78.3 | 3 | 0.14 | 2.94 | 21.74 |
| Q9QXZ6 | Solute carrier organican ion transporter family member/ Slco1a1 | 74.40 | 32.1 | 16 | 0.11 | 5.21 | 45.54 |
| P11588 | Major urinary protein 1/ Mup1 | 20.65 | 81.1 | 3 | 0.09 | 5.46 | 63.4 |
| Q9D154 | Leukocyte elastase inhibitor A/ Serpinb1a | 42.58 | 35.4 | 12 | 0.08 | 0.26 | 3.11 |
| Proteins accession | Proteins description | Gene | Peptides | Ratio | ||
|---|---|---|---|---|---|---|
| M/C | P/C | P/M | ||||
| Q63880 | Carboxylesterase 3A | Ces3a | LGIFGFLSTGDK | 0.15 | 3.08 | 19.87 |
| Q05816 | Fatty acid-binding protein 5 | Fabp5 | ELGVGLALR | 0.21 | 0.54 | 2.58 |
| Q9D154 | Leukocyte elastase inhibitor A | Serpinb1a | FQSLNAEVSK | 0.21 | 0.37 | 1.78 |
| Q9QXG4 | Acetyl-coenzyme A synthetase | Acss2 | TACPGPFLQYNFDVTK | 0.21 | 0.33 | 1.60 |
| Q9QXZ6 | Solute carrier organicanion transporter family member | Slco1a1 | GVQHPLYGEK | 0.44 | 10.53 | 23.69 |
| Q9DCY0 | Glycine N-acyltransferase-like protein | Keg1 | VIESLGATNLGK | 0.51 | 3.16 | 6.20 |
| G3X982 | Aldehyde oxidase 3 | Aox3 | TTWIAPGTLNDLLELK | 0.56 | 3.80 | 6.79 |
| Q99P30 | Peroxisomal coenzyme A | Nudt7 | EVFFVPLDYFLHPQVYYQK | 0.56 | 3.62 | 6.45 |
| Q9D7S7 | 60S ribosomal protein L22-like 1 | Rpl22l1 | TGNLGNVVHIER | 12.31 | 3.16 | 0.26 |
| P43276 | Histone H1.5 | H1-5 | GGVSLPALK | 8.06 | 2.71 | 0.34 |
| Q9D1M7 | Peptidyl-prolyl cis-trans isomerase | Fkbp11 | DPLVIELGQK | 6.74 | 2.34 | 0.35 |
Table 3 Differentially co-expressed proteins confirmed by targeted quantitative proteomics based on mass spectrometry
| Proteins accession | Proteins description | Gene | Peptides | Ratio | ||
|---|---|---|---|---|---|---|
| M/C | P/C | P/M | ||||
| Q63880 | Carboxylesterase 3A | Ces3a | LGIFGFLSTGDK | 0.15 | 3.08 | 19.87 |
| Q05816 | Fatty acid-binding protein 5 | Fabp5 | ELGVGLALR | 0.21 | 0.54 | 2.58 |
| Q9D154 | Leukocyte elastase inhibitor A | Serpinb1a | FQSLNAEVSK | 0.21 | 0.37 | 1.78 |
| Q9QXG4 | Acetyl-coenzyme A synthetase | Acss2 | TACPGPFLQYNFDVTK | 0.21 | 0.33 | 1.60 |
| Q9QXZ6 | Solute carrier organicanion transporter family member | Slco1a1 | GVQHPLYGEK | 0.44 | 10.53 | 23.69 |
| Q9DCY0 | Glycine N-acyltransferase-like protein | Keg1 | VIESLGATNLGK | 0.51 | 3.16 | 6.20 |
| G3X982 | Aldehyde oxidase 3 | Aox3 | TTWIAPGTLNDLLELK | 0.56 | 3.80 | 6.79 |
| Q99P30 | Peroxisomal coenzyme A | Nudt7 | EVFFVPLDYFLHPQVYYQK | 0.56 | 3.62 | 6.45 |
| Q9D7S7 | 60S ribosomal protein L22-like 1 | Rpl22l1 | TGNLGNVVHIER | 12.31 | 3.16 | 0.26 |
| P43276 | Histone H1.5 | H1-5 | GGVSLPALK | 8.06 | 2.71 | 0.34 |
| Q9D1M7 | Peptidyl-prolyl cis-trans isomerase | Fkbp11 | DPLVIELGQK | 6.74 | 2.34 | 0.35 |
| 1. |
Seitz HK, Bataller R, Cortez-Pinto H, et al. Alcoholic liver disease. Nat Rev Dis Primers 2018; 4: 16-38.
DOI PMID |
| 2. |
Zeng T, Zhang CL, Zhao N, et al. Impairment of Akt activity by CYP2E1 mediated oxidative stress is involved in chronic ethanol-induced fatty liver. Redox Biol 2018; 14: 295-304.
DOI PMID |
| 3. |
Singh S, Osna NA, Kharbanda KK. Treatment options for alcoholic and non-alcoholic fatty liver disease: A review. World J Gastroenterol 2017; 23: 6549-70.
DOI URL |
| 4. | Ding QX, Wang DX, Zhang XH. Research progress of Traditional Chinese Medicine in treatment of alcoholic fatty liver. Grassroots Medical Forum 2017; 21: 2124-26. |
| 5. | Wu XL, Feng X, Hao J, Yi X. Study on anti-alcoholic fatty liver effect of Gehua Jiejiu Dizhi decoction based on network pharmacology. Anhui Yi Ke Da Xue Xue Bao 2022; 57: 260-4. |
| 6. | Yi X, You SW, Long Y, Wang SS, Lu DM. Intervention Effect of Gehua Jiejiu Dizhi decoction on alcoholic fatty liver and liver PXR, CAR expression in mice. Zhong Guo Shi Yan Fang Ji Xue Za Zhi 2016; 22: 134-9. |
| 7. | Yi X, You SW, Long Y, Wang SS, Lu DM. Effect of Gehua Jiejue Dizhi decoction on the liver fatty deposition and expression of PXR in the mouse alcoholic fatty liver. Zhong Guo Bi Jiao Yi Xue Za Zhi 2017; 27: 36-42. |
| 8. | Yi X, Wang SS, Hao J, You SW, Zhan YM. Detection of LXRα-SREBP -1c signaling expression in liver of alcoholic fatty liver mice and intervention of Gehua Jiejiu Dizhi decoction. Zhong Guo Shi Yan Fang Ji Xue Za Zhi 2021; 39: 13-6+261. |
| 9. |
Li X, Wang W, Chen J. Recent progress in mass spectrometry proteomics for biomedical research. Sci China Life Sci 2017; 60: 1093-113.
DOI PMID |
| 10. |
Liu S, Li Z, Yu B, Wang S, Shen Y, Cong H. Recent advances on protein separation and purification methods. Adv Colloid Interface Sci 2020; 284: 102254-64.
DOI URL |
| 11. |
Yang L, Yang C, Thomes PG, et al. Lipophagy and alcohol-induced fatty liver. Front Pharmacol 2019; 10: 495-508.
DOI PMID |
| 12. |
Banerjee A, Russell WK, Jayaraman A, Ramaiah SK. Identification of proteins to predict the molecular basis for the observed gender susceptibility in a rat model of alcoholic steatohepatitis by 2-D gel proteomics. Proteomics 2008; 8: 4327-37.
DOI PMID |
| 13. |
Fernando H, Wiktorowicz JE, Soman KV, Kaphalia BS, Khan MF, Shakeel Ansari GA. Liver proteomics in progressive alcoholic steatosis. Toxicol Appl Pharmacol 2013; 266: 470-80.
DOI URL |
| 14. |
Klouckova I, Hrncirova P, Mechref Y, et al. Changes in liver protein abundance in inbred lcohol-preferring rats due to chronic alcohol exposure, as measured through a proteomics approach. Proteomics 2006; 6: 3060-74.
PMID |
| 15. |
Swart PC, Russell VA, Vlok NM, Dimatelis JJ. Early-ethanol exposure induced region-specific changes in metabolic proteins in the rat brain: a proteomics study. J Mol Neurosci 2018; 65: 277-88.
DOI PMID |
| 16. |
Aebersold R, Matthias M. Mass spectrometry-based proteomics. Nature 2003; 422: 198-207.
DOI |
| 17. |
Song P, Shen X. Proteomic analysis of liver in diet-induced Hyperlipidemic mice under Fructus Rosa roxburghii action. J Proteomics 2021; 230: 103982-94.
DOI URL |
| 18. | Dai J, Liang K, Zhao S, et al. Chemoproteomics reveals baicalin activates hepatic CPT1 to ameliorate diet-induced obesity and hepatic steatosis. Proc Natl Acad Sci USA 2018; 115: E5896-905. |
| 19. |
Du Y, Cai T, Li T, et al. Lysine malonylation is elevated in type 2 diabetic mouse models and enriched in metabolic associated proteins. Mol Cell Proteomics 2015; 14: 227-36.
DOI PMID |
| 20. |
Liu B, Jiang S, Li M, et al. Proteome-wide analysis of USP14 substrates revealed its role in hepatosteatosis via stabilization of FASN. Nat Commun 2018; 9: 4770-82.
DOI PMID |
| 21. |
Bertola A, Mathews S, Ki SH, Wang H, Gao B. Mouse model of chronic and binge ethanol feeding (the NIAAA model). Nat Protoc 2013; 8: 627-37.
DOI PMID |
| 22. | You M, Jogasuria A, Taylor C, Wu J. Sirtuin 1 signaling and alcoholic fatty liver disease. Hepatobiliary Surg Nutr 2015; 4: 88-100. |
| 23. |
Ren R, Wang Z, Wu M, Wang H. Emerging roles of SIRT1 in alcoholic liver disease. Int J Biol Sci 2020; 16: 3174-83.
DOI PMID |
| 24. | He PY, Hou ZP, Song CJ, Wang CQ, Li BQ, Mustapha UI. Resveratrol ameliorates experimental alcoholic liver disease by modulating oxidative stress. Evid Based Complement Alternat Med 2017; 2017: 4287890-901. |
| 25. |
Tang L, Yang F, Fang Z, Hu C. Resveratrol ameliorates alcoholic fatty liver by inducing autophagy. Am J Chin Med 2016; 44: 1207-20.
DOI URL |
| 26. |
Chen WM, Shaw LH, Chang PJ, et al. Hepatoprotective effect of resveratrol against ethanol-induced oxidative stress through induction of superoxide dismutase in vivo and in vitro. Exp Ther Med 2016; 11: 1231-8.
DOI URL |
| 27. |
McCormack L, Dutkowski P, El-Badry AM, Clavien PA. Liver transplantation using fatty livers: always feasible. J Hepatol 2011; 54: 1055-62.
DOI PMID |
| 28. |
Reis H, Peterek PT, Wohlschlaeger J, et al. Oil Red O-assessed macrosteatosis in liver transplantdonor biopsies predicts ischemia-reperfusion injury and clinical outcome. Virchows Arch 2014; 464: 165-74.
DOI URL |
| 29. |
Selzner M, Clavien PA. Fatty liver in liver transplantation and surgery. Semin Liver Dis 2001; 21: 105-13.
DOI PMID |
| 30. |
Sun XZ, Wang D, Ding L, et al. Activation of autophagy through the NLRP3/mTOR pathway: a potential mechanism for alleviation of pneumonia by Qingfei Yin. Front Pharmacol 2021; 12: 763160-77.
DOI URL |
| 31. |
Li X, Wu D, Niu J, et al. Intestinal Flora: a pivotal role in investigation of Traditional Chinese Medicine. Am J Chin Med 2021; 49: 237-68.
DOI URL |
| 32. |
Liang B, Gu N. Traditional Chinese Medicine for coronary artery disease treatment: clinical evidence from randomized controlled trials. Front Cardiovasc Med 2021; 8: 702110-9.
DOI URL |
| 33. | Yao ZS, Wu Y, Zhao HX, et al. Treatment of alcoholic liver disease with TCM. Heman Zhong Yi Yao Xue Bao 2020; 40: 1941-4. |
| 34. | Elken H, Xie JR, Liang GY, et al. Research progress of Traditional Chinese Medicine in the treatment of fatty liver disease. Liaoning Zhong Yi Yao Da Xue Xue Bao 2020; 22: 92-5. |
| 35. |
Ding XW, Xu YF, Nie P, et al. Changes in the serum metabolomic profiles of subjects with NAFLD in response to n-3 PUFAs and phytosterol ester: a double-blind randomized controlled trial. Food funct 2022; 13: 5189-201.
DOI URL |
| 36. |
Zhu S, Zhang J, Zhu D, et al. Adipose tissue plays a major role in retinoic acid-mediated metabolic homoeostasis. Adipocyte 2022; 11: 47-55.
DOI PMID |
| 37. |
Errafii K, Al-Akl NS, Khalifa O, Arredouani A. Comprehensive analysis of LncRNAs expression profiles in an in vitro model of steatosis treated with Exendin-4. J Transl Med 2021; 19: 235-46.
DOI |
| 38. |
Lian J, Nelson R, Lehner R. Carboxylesterases in lipid metabolism: from mouse to human. Protein Cell 2018; 9: 178-95.
DOI PMID |
| 39. |
Dominguez E, Galmozzi A, Chang JW, et al. Integrated phenotypic and activity-based profiling links Ces3 to obesity and diabetes. Nat Chem Biol 2014; 10: 113-21.
DOI PMID |
| 40. |
Mukherjee S, Choi M, Yun JW. Novel regulatory roles of carboxylesterase 3 in lipid metabolism and browning in 3T3-L1 white adipocytes. Appl Physiol Nutr Metab 2019; 44: 1089-98.
DOI PMID |
| 41. |
Yang L, Li X, Tang H, Gao Z, Zhang K, Sun K. A unique role of carboxylesterase 3 (Ces3) in beta-adrenergic signaling-stimulated thermogenesis. Diabetes 2019; 68: 1178-96.
DOI PMID |
| 42. |
Chiang JYL, Ferrell JM. Bile acid metabolism in liver pathobiology. Gene Expr 2018; 18: 71-87.
DOI PMID |
| 43. |
Zhang Y, Limaye PB, Lehman-McKeeman LD, Klaassen CD. Dysfunction of organic anion transporting polypeptide 1a1 alters intestinal bacteria and bile acid metabolism in mice. PLoS One 2012; 7: e34522-33.
DOI URL |
| 44. |
Zhang T, Li J, Liu CP, et al. Butyrate ameliorates alcoholic fatty liver disease via reducing endotoxemia an inhibiting liver gasdermin D-mediated pyroptosis. Ann Transl Med 2021; 9: 873-87.
DOI PMID |
| 45. |
Zhang Y, Csanaky IL, Cheng X, Lehman-McKeeman LD, Klaassen CD. Organic anion transporting polypeptide 1a1 null mice are sensitiveto cholestatic liver injury. Toxicol Sci 2012; 127: 451-62.
DOI URL |
| 46. |
Hartmann P, Hochrath K, Horvath A, et al. Modulation of the intestinal bile acid/farnesoid X receptor/fibroblast growth factor 15 axis improves alcoholic liver disease in mice. Hepatology (Baltimore, Md) 2018; 67: 2150-66.
DOI URL |
| 47. |
Jiang L, Chu HK, Gao B, et al. Transcriptomic profiling identifies novel hepatic and intestinal genes following chronic plus binge ethanol feeding in mice. Dig Dis Sci 2020; 65: 3592-604.
DOI |
| 48. |
Moore SM, Holt VV, Malpass LR, Hines IN, Wheeler MD. Fatty acid-binding protein 5 limits the anti-inflammatory response in murine macrophages. Mol Immunol 2015; 67: 265-75.
DOI PMID |
| 49. |
Babaev VR, Runner RP, Fan D, et al. Macrophage Mal1 deficiency suppresses atherosclerosis in low-density lipoprotein receptor-null mice by activating peroxisome proliferator-activated receptor-gamma-regulated genes. Arterioscler Thromb Vasc Biol 2011; 31: 1283-90.
DOI URL |
| 50. |
Matsuo M, Terai K, Kameda N, et al. Designation of enzyme activity of glycine-N-acyltransferase family genes and depression of glycine-N-acyltransferase in human hepatocellular carcinoma. Biochem Biophys Res Commun 2012; 420: 901-6.
DOI URL |
| 51. |
Shumar SA, Kerr EW, Geldenhuys WJ, et al. Nudt19 is a renal CoA diphosphohydrolase with biochemical and regulatory properties that are distinct from the hepatic Nudt7 isoform. J Biol Chem 2018; 293: 4134-48.
DOI PMID |
| 52. |
Shumar SA, Kerr EW, Fagone P, Infante AM, Leonardi R. Overexpression of Nudt7 decreases bile acid levels and peroxisomal fatty acidoxidation in the liver. J Lipid Res 2019; 60: 1005-19.
DOI URL |
| 53. |
Naquet P, Kerr EW, Vickers SD, Leonardi R. Leonardi R. Regulation of coenzyme A levels by degradation: the 'Ins and Outs'. Prog Lipid Res 2020; 78: 101028-72.
DOI URL |
| 54. |
Gasmi L, McLennan AG. The mouse Nudt7 gene encodes a peroxisomal nudix hydrolase specific for coenzyme A and its derivatives. Biochem J 2001; 357: 33-8.
DOI PMID |
| 55. |
Leonardi R, Rock CO, Jackowski S. Pank1 deletion in leptin-deficient mice reduces hyperglycaemia and hyperinsulinaemia and modifies global metabolism without affecting insulin resistance. Diabetologia 2014; 57: 1466-75.
DOI PMID |
| 56. |
Leonardi R, Rehg JE, Rock CO, Jackowski S. Pantothenate kinase 1 is required to support the metabolic transition from the fed to the fasted state. PLoS One 2010; 5: e11107-19.
DOI URL |
| 57. |
Kryvalap Y, Czyzyk J. The Role of proteases and serpin protease inhibitors in β-cell biology and diabetes. Biomolecules 2022; 12: 67-77.
DOI URL |
| 58. |
Liang X, Su Y, Huo Y. Forkhead box protein O1 (FoxO1) /SERPINB 1 ameliorates ROS production in diabetic nephropathy. Food Sci Nutr 2020; 9: 44-51.
DOI URL |
| 59. |
Xing YQ, Li A, Yang Y, Li XX, Zhang LN, Guo HC. The regulation of FOXO1 and its role in disease progression. Life Sci 2018; 193: 124-131.
DOI PMID |
| 60. |
Chen J, Lu Y, Tian M, Huang Q. Molecular mechanisms of FOXO1 in adipocyte differentiation. J Mol Endocrinol 2019; 62: R239-53.
DOI |
| 61. |
Skat-Rørdam J, Højland Ipsen D, Lykkesfeldt J, Tveden-Nyborg P. A role of peroxisome proliferator-activated receptor γ in non-alcoholic fatty liver disease. Basic Clin Pharmacol Toxicol 2019; 124: 528-37.
DOI URL |
| 62. | Huang Z, Zhang M, Plec AA, et al. ACSS2 promotes systemic fat storage and utilization through selective regulation of genes involved in lipid metabolism. Proc Natl Acad Sci USA 2018; 115: E9499-506. |
| 63. |
Martínez-Micaelo N, González-Abuín N, Terra X, et al. Identification of a nutrient-sensing transcriptional network in monocytes by using inbred rat models on a cafeteria diet. Dis Model Mech 2016; 9: 1231-9.
PMID |
| 64. |
Liu J. Ethanol and liver: recent insights into the mechanisms of ethanol-induced fatty liver. World J Gastroenterol 2014; 20: 14672-85.
DOI URL |
| 65. |
Terao M, Garattini E, Romão MJ, Leimkühler S. Evolution, expression, and substrate specificities of aldehyde oxidase enzymes in eukaryotes. J Biol Chem 2020; 295: 5377-89.
DOI PMID |
| 66. |
Zhong G, Seaman CJ, Paragas EM, et al. Aldehyde oxidase contributes to all-trans-retinoic acid biosynthesis in human liver. Drug Metab Dispos 2021; 49: 202-11.
DOI PMID |
| 67. | Zarei L, Farhad N, Abbasi A. All-trans retinoic acid (atRA) effectively improves liver steatosis in a rabbit model of high fat induced liver steatosis. Arch Physiol Biochem 2020: 1-6. |
| 68. |
Tsuchiya H. Retinoids as promising treatment for non-alcoholic fatty liver disease. Yakugaku Zasshi 2012; 132: 903-9.
DOI URL |
| 69. |
Lin IY, Yen CH, Liao YJ, et al. Identification of FKBP11 as a biomarker for hepatocellular carcinoma. Anticancer Research 2013; 33: 2763-9.
PMID |
| 70. | Momeni M, Kalir T, Farag S, Chuang L, Fishman D, Burstein DE. Expression of H1.5 and PLZF in granulosa cell tumors and normal ovarian tissues: a short report. Cellular Oncology (Dordrecht) 2014; 37: 229-34. |
| 71. |
Rao S, Peri S, Hoffmann J, et al. RPL22L 1 induction in colorectal cancer is associated with poor prognosis and 5-FU resistance. PLoS One 2019; 14: e0222392-412.
DOI URL |
| No related articles found! |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||

