Journal of Traditional Chinese Medicine ›› 2024, Vol. 44 ›› Issue (3): 505-514.DOI: 10.19852/j.cnki.jtcm.20240308.002
• Original articles • Previous Articles Next Articles
Identifying the geographical origin and processing technology of Moyao (Myrrh) on the basis of near-infrared spectroscopy combined with chemometrics
XU Ningning1, YAN Ganming2, XU Fengjie3, DENG Linfeng4, QIAO Xinjiang2, LU Changzheng5, CHENG Shaomin1( )
)
- 1 TCM Diagnosis Institute, College of Traditional Chinese Medicine, Jiangxi University of Chinese Medicine, Nanchang 330004, China
2 TCM Processing Institute, Pharmaceutical College, Jiangxi University of Chinese medicine, Nanchang 330004, China
3 School of Electronic Information and Artificial Intelligence, Shaanxi University of Science and Technology, Xi'an 710021, China
4 Jiangzhong Pharmaceutical Co., Ltd., Nanchang 330096, China
5 Jiangxi Guhan Refined Chinese Herbal Pieces Co., Ltd., Nanchang 330041, China
-
Received:2023-02-22Accepted:2023-05-18Online:2024-06-15Published:2024-03-08 -
Contact:Prof. CHENG Shaomin, College of Traditional Chinese Medicine, University of Chinese Medicine, Nanchang, Jiangxi 330004, China.csm21cn@139.com Telephone: +86-18079116281 -
Supported by:Jiangxi Provincial Administration of Traditional Chinese Medicine Key Research Laboratory on the Fundamentals of Chinese Medicine Evidence(Gan TCM Science and Education Word [2022] No. 8-4);Jiangxi University of Chinese Medicine Science and Technology Innovation Team Development Program: Traditional Chinese Medicine Constitution-State Identification Health Management Research Team(CXTD22016)
Cite this article
XU Ningning, YAN Ganming, XU Fengjie, DENG Linfeng, QIAO Xinjiang, LU Changzheng, CHENG Shaomin. Identifying the geographical origin and processing technology of Moyao (Myrrh) on the basis of near-infrared spectroscopy combined with chemometrics[J]. Journal of Traditional Chinese Medicine, 2024, 44(3): 505-514.
share this article
| Method | Group | Numbers | Total | |
|---|---|---|---|---|
| Training set | Prediction set | |||
| Origin | Kenya | 71 | 19 | 90 |
| Ethiopia | 62 | 28 | 90 | |
| Somalia | 69 | 21 | 90 | |
| Total | 202 | 68 | 270 | |
| Processing technology | Raw Moyao (Myrrh) | 62 | 28 | 90 |
| sprayed vinegar Moyao (Myrrh) | 75 | 15 | 90 | |
| soaked vinegar Moyao (Myrrh) | 65 | 25 | 90 | |
| Total | 202 | 68 | 270 | |
Table 1 Dataset of Moyao (Myrrh) samples, split into the training set and the prediction set
| Method | Group | Numbers | Total | |
|---|---|---|---|---|
| Training set | Prediction set | |||
| Origin | Kenya | 71 | 19 | 90 |
| Ethiopia | 62 | 28 | 90 | |
| Somalia | 69 | 21 | 90 | |
| Total | 202 | 68 | 270 | |
| Processing technology | Raw Moyao (Myrrh) | 62 | 28 | 90 |
| sprayed vinegar Moyao (Myrrh) | 75 | 15 | 90 | |
| soaked vinegar Moyao (Myrrh) | 65 | 25 | 90 | |
| Total | 202 | 68 | 270 | |
| Method | Preprocessing | Precision | Accuracy of training test | Accuracy of test | F1score | Recall |
|---|---|---|---|---|---|---|
| KNN | Raw | 0.8889 | 1 | 0.8971 | 0.8987 | 0.8935 |
| SNV | 0.9833 | 1 | 0.9853 | 0.9853 | 0.9841 | |
| MSC | 0.9833 | 1 | 0.9853 | 0.9853 | 0.9841 | |
| FD | 0.9683 | 1 | 0.9706 | 0.9706 | 0.9683 | |
| SD | 0.9683 | 1 | 0.9706 | 0.9706 | 0.9683 | |
| SNV+FD | 0.9666 | 1 | 0.9706 | 0.9706 | 0.9666 | |
| SVM | Raw | 0.8929 | 0.9109 | 0.8676 | 0.8663 | 0.8611 |
| SNV | 0.9545 | 0.9703 | 0.9559 | 0.9558 | 0.9524 | |
| MSC | 0.9545 | 0.9703 | 0.9559 | 0.9558 | 0.9524 | |
| FD | 0.9683 | 0.995 | 0.9706 | 0.9706 | 0.9683 | |
| SD | 0.9683 | 0.995 | 0.9706 | 0.9706 | 0.9683 | |
| SNV+FD | 0.9683 | 0.9901 | 0.9706 | 0.9706 | 0.9683 | |
| PLS-DA | Raw | 0.9666 | 0.9208 | 0.9706 | 0.9706 | 0.9666 |
| MSC | 0.9833 | 0.9752 | 0.9853 | 0.9853 | 0.9841 | |
| SNV | 0.971 | 0.9356 | 0.9706 | 0.9705 | 0.9666 | |
| FD | 0.9833 | 0.9653 | 0.9853 | 0.9853 | 0.9841 | |
| SD | 0.9833 | 0.9752 | 0.9853 | 0.9853 | 0.9841 | |
| SNV+FD | 0.9683 | 0.9604 | 0.9706 | 0.9706 | 0.9683 |
Table 2 Precision, accuracy, F1 score, and recall (%) of the KNN, SVM, and PLS-DA classification methods for Moyao (Myrrh) from different geographical origins
| Method | Preprocessing | Precision | Accuracy of training test | Accuracy of test | F1score | Recall |
|---|---|---|---|---|---|---|
| KNN | Raw | 0.8889 | 1 | 0.8971 | 0.8987 | 0.8935 |
| SNV | 0.9833 | 1 | 0.9853 | 0.9853 | 0.9841 | |
| MSC | 0.9833 | 1 | 0.9853 | 0.9853 | 0.9841 | |
| FD | 0.9683 | 1 | 0.9706 | 0.9706 | 0.9683 | |
| SD | 0.9683 | 1 | 0.9706 | 0.9706 | 0.9683 | |
| SNV+FD | 0.9666 | 1 | 0.9706 | 0.9706 | 0.9666 | |
| SVM | Raw | 0.8929 | 0.9109 | 0.8676 | 0.8663 | 0.8611 |
| SNV | 0.9545 | 0.9703 | 0.9559 | 0.9558 | 0.9524 | |
| MSC | 0.9545 | 0.9703 | 0.9559 | 0.9558 | 0.9524 | |
| FD | 0.9683 | 0.995 | 0.9706 | 0.9706 | 0.9683 | |
| SD | 0.9683 | 0.995 | 0.9706 | 0.9706 | 0.9683 | |
| SNV+FD | 0.9683 | 0.9901 | 0.9706 | 0.9706 | 0.9683 | |
| PLS-DA | Raw | 0.9666 | 0.9208 | 0.9706 | 0.9706 | 0.9666 |
| MSC | 0.9833 | 0.9752 | 0.9853 | 0.9853 | 0.9841 | |
| SNV | 0.971 | 0.9356 | 0.9706 | 0.9705 | 0.9666 | |
| FD | 0.9833 | 0.9653 | 0.9853 | 0.9853 | 0.9841 | |
| SD | 0.9833 | 0.9752 | 0.9853 | 0.9853 | 0.9841 | |
| SNV+FD | 0.9683 | 0.9604 | 0.9706 | 0.9706 | 0.9683 |
| Model | Preprocessing | Precision | Acc_Train | Acc_Test | F1 score | Recall |
|---|---|---|---|---|---|---|
| KNN | Raw | 0.9122 | 1.0000 | 0.9118 | 0.9089 | 0.9200 |
| SNV | 0.8833 | 1.0000 | 0.8971 | 0.8987 | 0.8978 | |
| MSC | 0.9267 | 1.0000 | 0.9412 | 0.9418 | 0.9378 | |
| FD | 0.9753 | 1.0000 | 0.9706 | 0.9701 | 0.9556 | |
| SD | 0.9246 | 1.0000 | 0.9265 | 0.9245 | 0.8978 | |
| SNV+FD | 0.9444 | 1.0000 | 0.9559 | 0.9561 | 0.9511 | |
| SVM | Raw | 0.6205 | 0.7723 | 0.6324 | 0.6250 | 0.6383 |
| SNV | 0.9540 | 0.9406 | 0.9412 | 0.9388 | 0.9111 | |
| MSC | 0.9540 | 0.9307 | 0.9412 | 0.9388 | 0.9111 | |
| FD | 0.9505 | 0.9653 | 0.9559 | 0.9556 | 0.9422 | |
| SD | 0.9139 | 0.9802 | 0.9265 | 0.9259 | 0.9067 | |
| SNV+FD | 0.9139 | 0.9455 | 0.9265 | 0.9259 | 0.9067 | |
| PLS-DA | Raw | 0.8500 | 0.9109 | 0.8676 | 0.8697 | 0.8622 |
| SNV | 0.7491 | 0.8564 | 0.7647 | 0.7681 | 0.7643 | |
| MSC | 0.7750 | 0.8416 | 0.7941 | 0.7963 | 0.7895 | |
| FD | 0.7776 | 0.9010 | 0.7794 | 0.7722 | 0.8057 | |
| SD | 0.8136 | 0.8515 | 0.8088 | 0.8012 | 0.8192 | |
| SNV+FD | 0.8521 | 0.9257 | 0.8382 | 0.8382 | 0.8310 |
Table 3 Precision, accuracy, F1 score, and recall (%) of the KNN, SVM and PLS-DA classification methods for distinguishing the different processing technologies
| Model | Preprocessing | Precision | Acc_Train | Acc_Test | F1 score | Recall |
|---|---|---|---|---|---|---|
| KNN | Raw | 0.9122 | 1.0000 | 0.9118 | 0.9089 | 0.9200 |
| SNV | 0.8833 | 1.0000 | 0.8971 | 0.8987 | 0.8978 | |
| MSC | 0.9267 | 1.0000 | 0.9412 | 0.9418 | 0.9378 | |
| FD | 0.9753 | 1.0000 | 0.9706 | 0.9701 | 0.9556 | |
| SD | 0.9246 | 1.0000 | 0.9265 | 0.9245 | 0.8978 | |
| SNV+FD | 0.9444 | 1.0000 | 0.9559 | 0.9561 | 0.9511 | |
| SVM | Raw | 0.6205 | 0.7723 | 0.6324 | 0.6250 | 0.6383 |
| SNV | 0.9540 | 0.9406 | 0.9412 | 0.9388 | 0.9111 | |
| MSC | 0.9540 | 0.9307 | 0.9412 | 0.9388 | 0.9111 | |
| FD | 0.9505 | 0.9653 | 0.9559 | 0.9556 | 0.9422 | |
| SD | 0.9139 | 0.9802 | 0.9265 | 0.9259 | 0.9067 | |
| SNV+FD | 0.9139 | 0.9455 | 0.9265 | 0.9259 | 0.9067 | |
| PLS-DA | Raw | 0.8500 | 0.9109 | 0.8676 | 0.8697 | 0.8622 |
| SNV | 0.7491 | 0.8564 | 0.7647 | 0.7681 | 0.7643 | |
| MSC | 0.7750 | 0.8416 | 0.7941 | 0.7963 | 0.7895 | |
| FD | 0.7776 | 0.9010 | 0.7794 | 0.7722 | 0.8057 | |
| SD | 0.8136 | 0.8515 | 0.8088 | 0.8012 | 0.8192 | |
| SNV+FD | 0.8521 | 0.9257 | 0.8382 | 0.8382 | 0.8310 |

Figure 1 Spectra of all Moyao (Myrrh) samples in the wavelength range of 900-1600 nm A: original spectra; B: spectra after pretreatment by FD; C: spectra after pretreatment by SD; D: spectra after pretreatment by SNV; E: spectra after pretreatment by MSC; F: spectra after pretreatment by MSC + FD. FD: first derivation; SD: second derivation; SNV: standard normal variation; MSC: multiplicative signal correction; MSC: multiplicative signal correction.
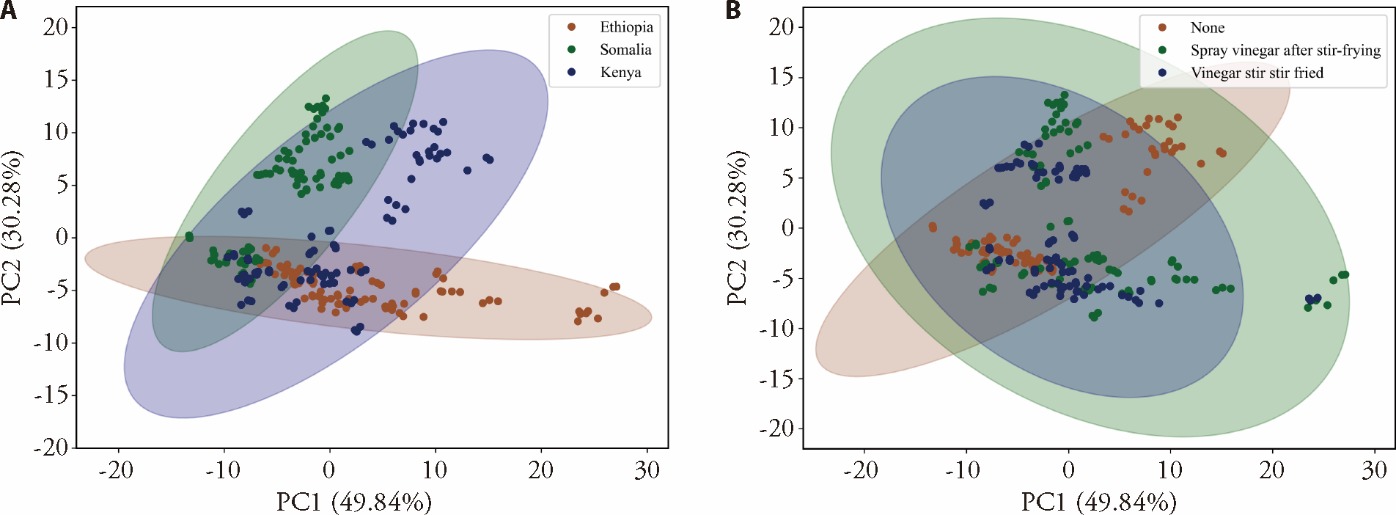
Figure 2 PCA scores plot in the spectral range of 908.1-1676.2 nm A: different geographical origins; B: different processing technologies. PCA: principal component analysis.
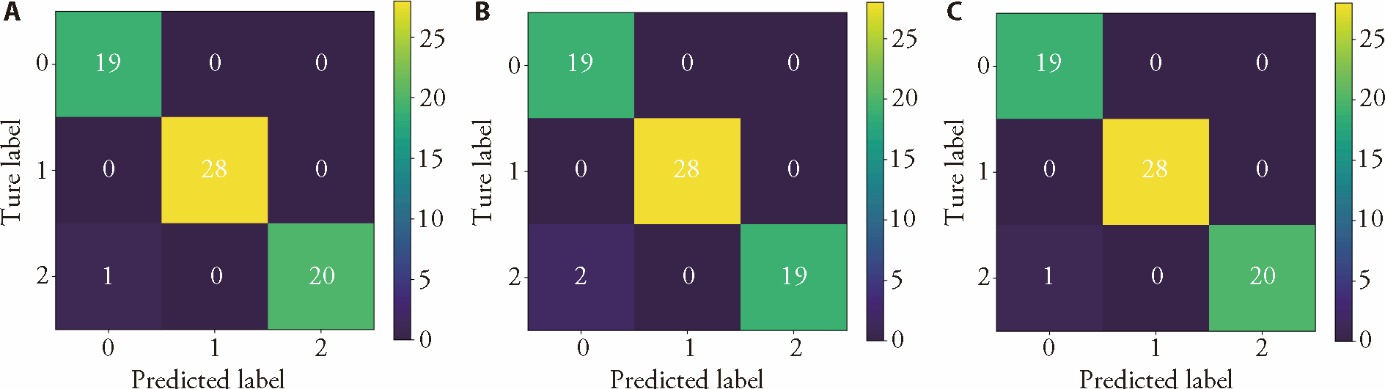
Figure 3 Confusion matrix of the geographic origin classification models A: confusion Matrix of k-nearest neighbor; B: confusion Matrix of support vector machine; C: confusion matrix of partial least squares classification analysis.
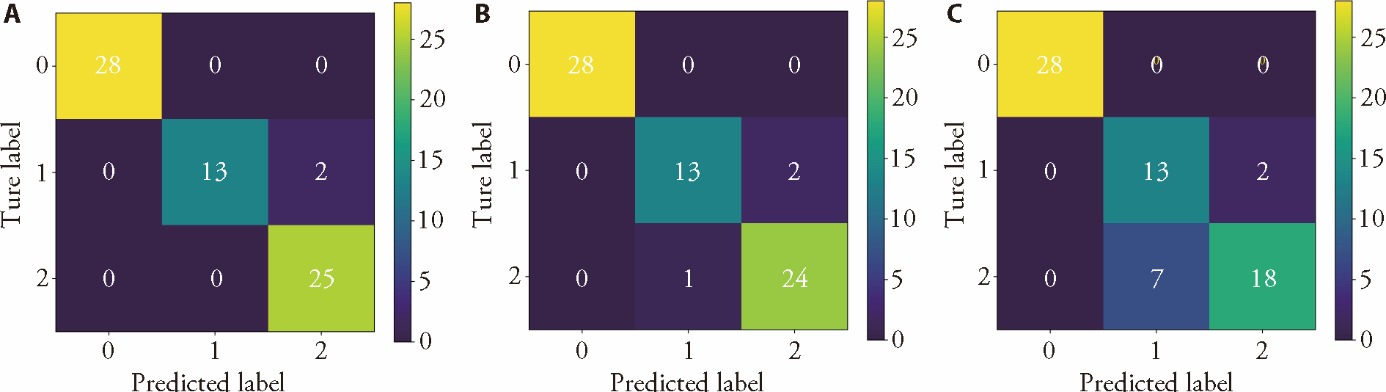
Figure 4 Confusion matrix of the processing technology classification models A: confusion Matrix of k-nearest neighbor; B: confusion Matrix of support vector machine; C: confusion matrix of partial least squares classification analysis.
| 1. | Han L, Sun JY, Zhou L, Fu XY, Bai CC. Research progress on non-drug chemical components and drug effects. Asian J Tradit Med 2015; 11: 38-42. |
| 2. | Dolara P, Luceri C, Ghelardini C, et al. Analgesic effects of myrrh. Nature 1996; 379: 29. |
| 3. | Zhen Q. Yao Xing Lun. Wuhu: research department of Wan'an Medical College, 1983: 42. |
| 4. | Wang XQ, C LL, Luo JQ, Liang SW. Historical evolution and modern research on myrrh processing. Ya Tai Chuan Tong Yi Yao 2016; 12: 66-9. |
| 5. |
Abbas RK, Al-Mushhin AAM, Elsharbasy FS, Ashiry KO. Nutritive value, polyphenol constituents and prevention of pathogenic microorganism by different resin extract of commiphora myrrh. J Pure Appl Microbiol 2020; 14: 1871-78.
DOI URL |
| 6. | Chinese Pharmacopoeia Commission. Pharmacopoeia of the People's Republic of China, Part 1. Beijing: China Medical Science Press, 2020: 193. |
| 7. | Jiangxi Food and Drug Administration. Preparation standard of TCM decoction pieces in Jiangxi province 2008 edition. Shanghai: Shanghai Science and Technology Press, 2009: 454. |
| 8. | Gong QF. A complete book of chinese medicine processing of Zhangshubang. Nanchang: Jiangxi Science and Technology Press, 1990: 384. |
| 9. | Yu XL, Sun L, Xu JM, Li G, Ma SC. The basic textual research of natural myrrh, colloidal myrrh and muku myrrh. Zhong Guo Yao Shi 2016; 30: 466 -71. |
| 10. | Li X, Wu MQ, Lin FJ, Chen P. Research on fingerprints of myrrha from different origins. Zhong Yi Yao Dao Bao 2019; 25: 50-4. |
| 11. | Lu JR, Li WB, Xu N, et al. Quality status analysis and intrinsic connection research of growing place, morphological characteristics, and quality of Chinese medicine: Cyperi Rhizoma (Xiangfu) as a case study. Evid Based Complement Alternat Med 2022. |
| 12. |
Qiu F, Wu S, Lu XR, et al. Quality evaluation of the artemisinin-producing plant Artemisia annua L. based on simultaneous quantification of artemisinin and six synergistic components and hierarchical cluster analysis. Ind Crops Prod 2018; 118: 131-41.
DOI URL |
| 13. | REN H, ZHAO LT, GAO K, et al. Deciphering the chemical profile and pharmacological mechanism of Jinlingzi powder against bile reflux gastritis using ultra-high performance liquid chromatography coupled with Q exactive focus mass spectrometry, network pharmacology, and molecular docking. J Tradit Chin Med 2023; 43: 1209-18. |
| 14. |
Wang Y, He T, Wang JJ, et al. High performance liquid chromatography fingerprint and headspace gas chromatography-mass spectrometry combined with chemometrics for the species authentication of Curcumae Rhizoma. J Pharm Biomed Anal 2021; 202: 114144.
DOI URL |
| 15. |
He Y, Wang JZ, Wang MZ, Zhang JF. Discrimination of wild and domestic deer musk using isotope ratio mass spectrometry. J Mass Spectrom 2018; 53: 1078-85.
DOI PMID |
| 16. | Li HY, Sun JY. Comparative study on principal components of essential oil of myrrh before and after processing by GC-MS. Zhongg Cheng Yao 1998; 20: 19-20. |
| 17. |
Qudsia T, Tahir M, Sibtain A, et al. Characterization and anticancer potential of Withania somnifera fruit bioactives (a native species to Pakistan) using gas chromatography-mass spectrometer, nuclear magnetic resonance and liquid chromatography-mass spectrometry-electrospray ionization. J Tradit Chin Med 2022; 42: 908-14.
DOI |
| 18. | Santhosh Kumar JU, Krishna V, Seethapathy GS, Ganesan R, Ravikanth G, Shaanker RU. Assessment of adulteration in raw herbal trade of important medicinal plants of India using DNA barcoding. Biotech 2018; 8: 135. |
| 19. | Zhong YC, Wang HY, Wei QH, et al. Combining DNA barcoding and HPLC fingerprints to trace species of an important Traditional Chinese Medicine fritillariae bulbus. Molecules 2019; 24. |
| 20. |
Wang Q, Wang XH, Wu XJ, et al. 1H NMR-based metabolic profiling approach to identify the geo-authentic Chinese yam (Dioscorea polystachya Turczaninow cv. Tiegun). J Food Compos Anal 2021; 98: 103805.
DOI URL |
| 21. | An L, Yuan Y, Ma J, et al. NMR-based metabolomics approach to investigate the distribution characteristics of metabolites in Dioscorea opposita Thunb. cv. Tiegun. Food Chem 2019; 298: 125063. |
| 22. | Pasquini C. Near infrared spectroscopy: a mature analytical technique with new perspectives - a review. Anal Chim Acta 2018: 1026, 8-36. |
| 23. |
Jamrogiewicz M. Application of the near-infrared spectroscopy in the pharmaceutical technology. J Pharm Biomed Anal 2012; 66: 1-10.
DOI PMID |
| 24. |
Blanco M, Gozalez Bano R, Bertran E. Monitoring powder blending in pharmaceutical processes by use of near infrared spectroscopy. Talanta 2002; 56: 203-12.
PMID |
| 25. |
Varrà MO, Fasolato L, Serva L, Ghidini S, Novelli E, Zanardi E. Use of near infrared spectroscopy coupled with chemometrics for fast detection of irradiated dry fermented sausages. Food Control 2020; 110: 107009.
DOI URL |
| 26. |
Xu Y, Zhang J, Wang Y. Recent trends of multi-source and non-destructive information for quality authentication of herbs and spices. Food Chem 2022; 398: 133939.
DOI URL |
| 27. |
Coppa M, Ferlay A, Leroux C, et al. Prediction of milk fatty acid composition by near infrared reflectance spectroscopy. Int Dairy J 2010; 20: 182-9.
DOI URL |
| 28. |
Zhou Y, Zuo Z, Xu F, Wang Y. Origin identification of Panax notoginseng by multi-sensor information fusion strategy of infrared spectra combined with random forest. Spectrochim Acta Part A 2020; 226: 117619.
DOI URL |
| 29. |
Manuel MNB, da Silva AC, Lopes GS, Ribeiro LPD. One-class classification of special agroforestry Brazilian coffee using NIR spectrometry and chemometric tools. Food Chem 2022; 366: 130480.
DOI URL |
| 30. |
Lan Z, Zhang Y, Sun Y, et al. Rapid quantitative detection of the discrepant compounds in differently processed Curcumae Rhizoma products by FT-NIR combined with VCPA-GA technology. J Pharm Biomed Anal 2021; 195: 113837.
DOI URL |
| 31. |
Aminu M, Ahmad NA. Complex chemical data classification and discrimination using locality preserving partial least squares discriminant analysis. ACS Omega 2020; 5: 26601-10.
DOI PMID |
| 32. |
Keller JM, Gray MR, Givens JA. A fuzzy K-nearest neighbor algorithm. IEEE Trans Syst Man Cybern 1985; 4: 580-5.
DOI URL |
| 33. |
de Santana FB, Otani SK, de Souza AM, Poppi RJ. Comparison of PLS and SVM models for soil organic matter and particle size using vis-NIR spectral libraries. Geoderma Regional 2021; 27: e00436.
DOI URL |
| 34. |
Brown SSG, Mak E, Clare I, et al. Support vector machine learning and diffusion-derived structural networks predict amyloid quantity and cognition in adults with Down's syndrome. Neurobiol Aging 2022; 115: 112-21.
DOI PMID |
| 35. |
Snee RD. Validation of regression models: methods and examples. Technometrics 1977; 19: 415-28.
DOI URL |
| 36. |
Wu W, Walczak B, Massart DL, et al. Artificial neural networks in classification of NIR spectral data: design of the training set. Chemom Intell Lab Syst 1996; 33: 35-46.
DOI URL |
| 37. |
Rajer-Kanduč K, Zupan J, Majcen N. Separation of data on the training and test set for modelling: a case study for modelling of five colour properties of a white pigment. Chemom Intell Lab Syst 2003; 65: 221-29.
DOI URL |
| 38. |
Galvao RK, Araujo MC, Jose GE, Pontes MJ, Silva EC, Saldanha TC. A method for calibration and validation subset partitioning. Talanta 2005; 67: 736-40.
DOI PMID |
| 39. |
Orlandi G, Calvini R, Pigani L, et al. Electronic eye for the prediction of parameters related to grape ripening. Talanta 2018; 186: 381-8.
DOI PMID |
| 40. | Ciocca G, Napoletano P, Schettini R. CNN-based features for retrieval and classification of food images. Comput Vis Imagge Und 2018; 176: 70-7. |
| No related articles found! |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||

